Precision medicine services
Precision medicine services
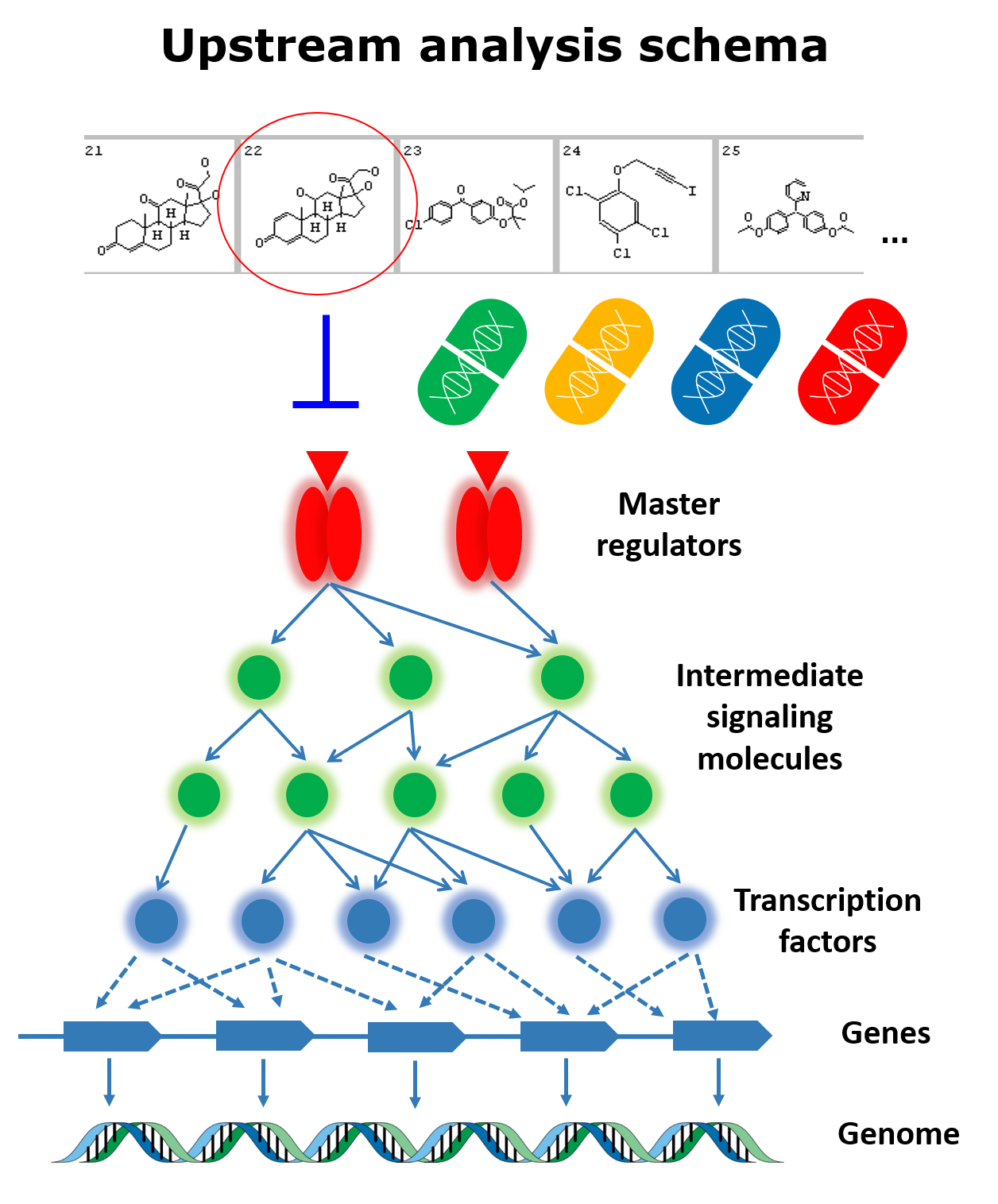
Our in-house omics data processing approaches powered by the manually collected for over 30 years knowledge bases on transcription regulation and signaling pathways allow us to offer unique service in the area of precision medicine: personalised reconstruction of the molecular mechanism of the pathology of a given patient based on the collected omics data. Our technology allows construction of a personalised model of the patient’s pathology, which is based on the Upstream Analysis approach and which identifies master-regulators responsible for development of the studied disease. We are capable to not only identify these master-regulators, but also identify chemical compounds, which include approved drugs and drugs undergoing clinical trials, as well as drug-like compounds, that can be effective in the studied case.
The analysis approach is dependent on the available omics data for the studied clinical case. We can work with genomics, proteomics, epigenomics, metabolomics, and transcriptomics patient data in any combinations (solely or by integrating various omics data types to fine-tune the constructed model). We can work with any disease, and you can view demo reports on analyses performed for such pathologies as cancer, neurodegenerative diseases, infectious diseases, diabetes and metabolic diseases, hypertension.
Our analysis approach identifies activated targets in the examined patient data by integrating promoter and enhancer analysis with reconstruction of a complex network of signal transduction pathways that are activated in the studied pathology. As a result, it identifies master-regulators of the studied disease and suggests known and re-purposing drugs that can be effective in the given clinical case. It is based on worldwide known and accepted databases: TRANSFAC®, TRANSPATH®, and HumanPSD™. The analysis can be launched either on raw or pre-processed patient omics data, As a result, we will provide a comprehensive report on the identified drug targets and prospective therapies.
In addition to our in-house approaches towards patient omics data analysis, for a number of cancer pathologies we can also provide a classical molecular tumor board report, matching the identified mutations in the patient’s pathology to the associated treatment outcomes. You can view an example of MTB report generated for colorectal cancer patient case here.
The precision medicine services we are providing are not licensed as a medical advice and should not be considered as such. The performed analysis is intended for research usage, however, we know that some medical doctors do consider the prospective treatments suggested by our models for their patients in complicated clinical cases when no conventionally accepted treatments are expected to give a positive treatment response. Our predictions are based purely on the reconstructed molecular mechanism of the patient’s pathology and there can be no warranties that any of the predicted drug agents will indeed be effective for the given patient.
You are welcome to explore our precision medicine analysis demo reports based on various omics data input types and different pathology origins:
- Colorectal Cancer (Personalized patient data) — Genomics, VCF
- MTB (Molecular Tumor Board) report example for colorectal cancer patient — Genomics, VCF
- Esophageal Squamous Cell Carcinoma (GSE32424) — Transcriptomics, FASTQ
- IFN-alpha induction (GSE31193) — Transcriptomics, LogFC Table
- Lung cancer, treatment by TGF (ST000010) — Metabolomics, Table
- Osteosarcoma, neoplasm metastasis (GSE66789) — Transcriptome + Proteome, RNA-seq + Mass-spec proteomics
- Ovarian cancer, cisplatin-resistance (GSE15709) — Transcriptomics + Epigenomics, CEL + BED
- SNP associated with Diabetes Mellitus — Genomics, SNP list
- Parkinson disease, induced a-Syn expression in SH-SY5Y cells (GSE145804) — Transcriptomics, LogFC Table
- Non-Small Cell Lung Carcinoma (NCI-H1975) — Genomics, VCF
- MTB (Molecular Tumor Board) report example for non-small cell lung carcinoma (NCI-H1975) — Genomics, VCF
- Hypertension (GSE157131) — Epigenomics, cg lists
Disclaimer
The results of analysis, contained in any of the reports produced by this service, are intended for research use only and should not be used for medical or professional advice. GeneXplain GmbH makes no guarantee of the comprehensiveness, reliability or accuracy of the information contained in the generated analysis reports.
Decisions regarding care and treatment of patients should be fully made by attending doctors. The predicted chemical compounds listed in the reports are given only for doctor’s consideration and they cannot be treated as prescribed medication. It is the physician’s responsibility to independently decide whether any, none or all of the predicted compounds can be used solely or in combination for patient treatment purposes, taking into account all applicable information regarding FDA prescribing recommendations for any therapeutic and the patient’s condition, including, but not limited to, the patient’s and family’s medical history, physical examinations, information from various diagnostic tests, and patient preferences in accordance with the current standard of care. Whether or not a particular patient will benefit from a selected therapy is based on many factors and can vary significantly.
The compounds predicted to be active against the identified drug targets in the reports are not guaranteed to be active against any particular patient’s condition. GeneXplain GmbH does not give any assurances or guarantees regarding the treatment information and conclusions given in the reports. There is no guarantee that any third party will provide a refund for any of the treatment decisions made based on these results. None of the listed compounds was checked by us for adverse side-effects or even toxic effects.
The analysis reports contain information about chemical drug compounds, clinical trials and disease biomarkers retrieved from the HumanPSD™ database of gene-disease assignments maintained and exclusively distributed worldwide by geneXplain GmbH. The information contained in this database is collected from scientific literature and public clinical trials resources. It is updated to the best of geneXplain’s knowledge however we do not guarantee completeness and reliability of this information leaving the final checkup and consideration of the predicted therapies to the medical doctor. In all cases, the end user (including researchers and medical doctors) accepts full responsibility for all risks associated with using of information, contained in the reports generated as a result of our service.
The scientific analysis underlying our service reports employs a complex analysis pipeline which uses geneXplain’s proprietary Upstream Analysis approach, integrated with TRANSFAC® and TRANSPATH® databases maintained and exclusively distributed worldwide by geneXplain GmbH. The pipeline and the databases are updated to the best of geneXplain’s knowledge and belief, however, geneXplain GmbH shall not give a warranty as to the characteristics or to the content and any of the results produced by our analysis methods. Moreover, any warranty concerning the completeness, up-to-dateness, correctness and usability of provided results shall be excluded.
The analysis results produced by us severely depend on the quality of input data used for the analysis.
Note that the text given in the reports is not unique and can be fully or partially repeated in our other analysis reports. This should be considered when publishing any results or excerpts from the provided reports. This restriction refers only to the general description of analysis methods used for generating the reports. All data and graphics referring to the concrete set of input data, including lists of mutated genes, differentially expressed genes/proteins/metabolites, functional classifications, identified transcription factors and master regulators, constructed molecular networks, lists of chemical compounds and reconstructed model of molecular mechanisms of the studied pathology are unique in respect to the used input data set used for the analysis launch.