Genome Enhancer
Genome Enhancer is available as part of the TRANSFAC Diseases package
Fully Automated One-Click Multi-Omics Discovery
From Raw Omics Data to Drug Targets and Therapies — in One Click
Genome Enhancer offers a fully automated, one-click solution for discovering disease mechanisms and identifying potential drug targets from patient-derived omics data. This pipeline eliminates the need for manual data preprocessing or complex scripting—empowering both bench scientists and clinicians to generate publication-ready insights from their data with just a few inputs.
Whether you’re working with transcriptomics, DNA methylation, or combined datasets, Genome Enhancer delivers a complete, interpretable report—including pathway diagrams, regulatory networks, transcription factor analyses, and prioritized drug candidates.
This solution is ideal for anyone seeking a fast, reproducible, and scientifically validated approach to extract biological meaning from omics data without deep bioinformatics expertise. Proven applications of Genome Enhancer include cancer, neurodegenerative diseases, infectious diseases, diabetes and metabolic diseases, hypertension.
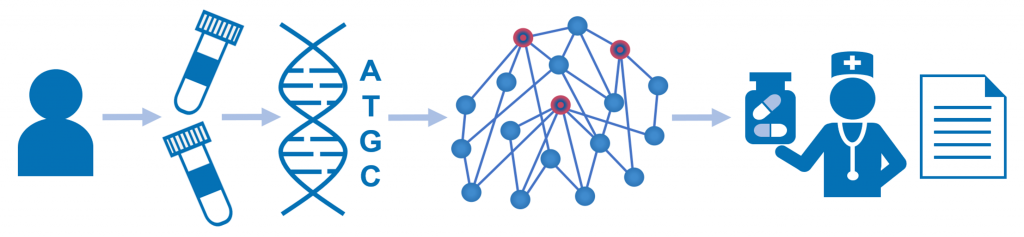
Genome Enhancer uses Upstream Analysis, an integrated promoter and pathway analysis, to identify potential drug targets of the studied pathology.
In the first step of this analysis the transcription factors that regulate differentially expressed or mutated genes are identified with the use of the TRANSFAC® database of transcription factors binding sites.
The second step searches for common master-regulators of the identified transcription factors by building a personalized signal transduction network of the studied pathology using the TRANSPATH® database of mammalian signal transduction and metabolic pathways. The identified
master regulators are prospective drug target candidates. They are used for further selection of chemical compounds that can bring therapeutic benefit for the studied clinical case. In this step the HumanPSD™database is employed to identify drugs that have been tested in clinical trials. The cheminformatic tool PASS predicts small molecules that can affect the identified targets.
Finally, Genome Enhancer generates a comprehensive analysis report about the personalized drug targets identified for a certain patient, or a group of patients, and the drugs that may be effective in this case. You can view a number of Genome Enhancer demo reports at the corresponding section of this page.
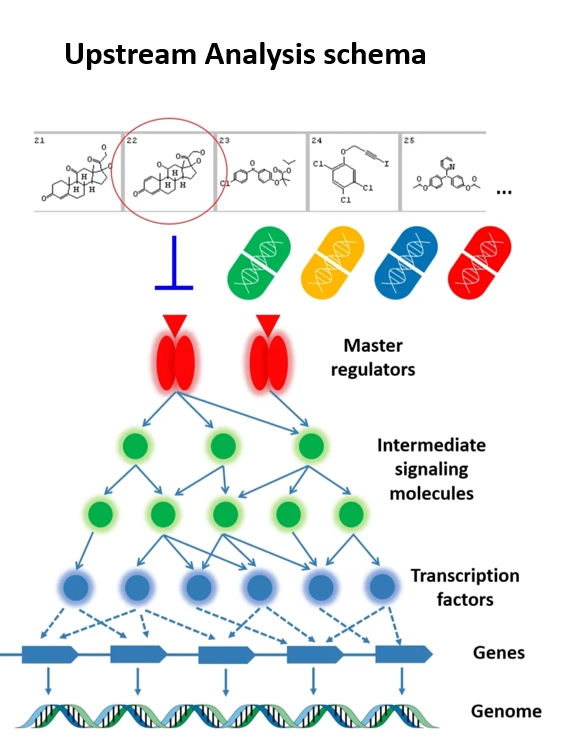
Key Features of the One-Click Genome Enhancer Workflow
Graphical Experiment Setup
No coding needed—just upload your omics files and define groups (e.g., control vs. disease) via a simple drag-and-drop interface. Choose between analysis modes (e.g., transcriptomics, epigenomics, or multi-omics).
Multi-omics integration
Genome Enhancer provides flexible integration of all five “-omics” data types:
Transcriptomics, Genomics, Epigenomics, Proteomics, Metabolomics.
Any combinations as well as individual omics data can be combined in one analysis run.
The omics integration is done following the principles of organisation of molecular-biological and biochemical systems in eukaryotic cells. The Upstream Analysis integrates promoter and pathway analysis, to identify potential drug targets of the studied pathology. Transcriptomics data help to find differentially expressed genes (DEGs); Metabolomics help to reveal which of the DEGs are most critical for metabolome changes in the studied pathology; Epigenomic data help to identify most regulatory active genomic regions for searching for TFBS enrichment; Genomic data help to reveal TF binding sites affected by regulatory mutations; Proteomics data help to strengthen the master regulator search.
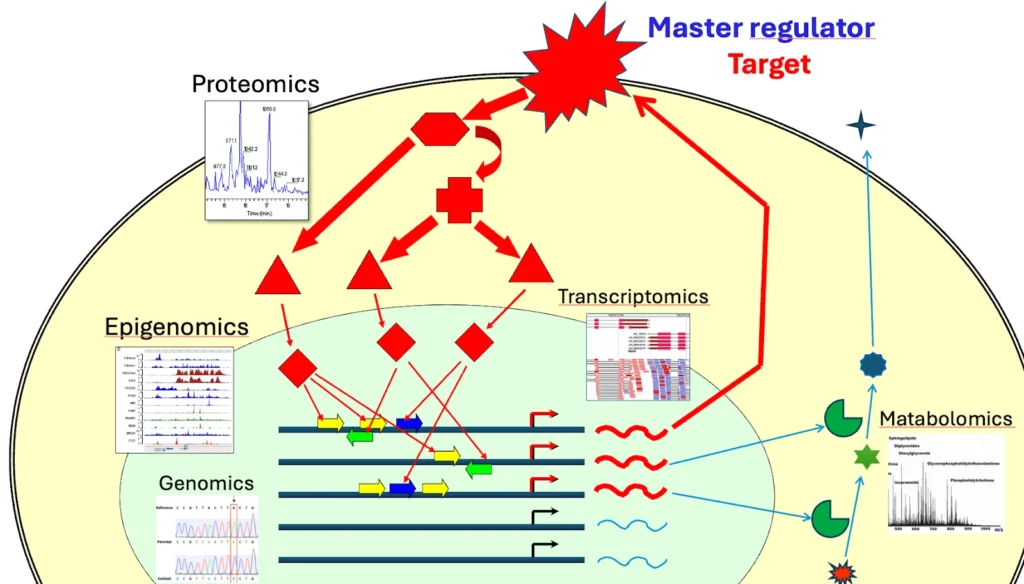
By integrating different omics data, researchers can move beyond correlations to causal insights – for example, identifying master regulator genes or proteins that when perturbed rewire downstream pathways and drive a disease phenotype .
In short, multi-omics integration helps to identify the key drivers (“molecular mechanisms”) of a condition that would be missed by single-omics approach. In practical terms, a multi-omics strategy can show how changes at one level (say DNA methylation in the epigenome) can lead to changes in gene expression in the transcriptome which in turn affect protein signaling networks.
This holistic view is key for complex issues like drug resistance in cancer where changes in gene regulation, signaling pathways and even metabolism can all intertwine. By looking at multiple layers together, scientists can identify critical points in the system – for example a transcription factor or receptor that if targeted could modulate the activity of an entire network of disease related genes . This systems-level view is what makes multi-omics integration so powerful in modern biomedical research.
Key Steps in Multi-Omics Analysis for Pathway Discovery
Pathway discovery from multi-omics data is a multi-step process that integrates diverse layers of molecular information. While several open-source and commercial tools can support each step, the overall process follows a logical structure:
Step 1: Preprocessing and Differential Analysis
Researchers begin by performing quality control, normalization, and statistical tests to identify significant molecular differences across conditions—typically genes, methylation sites, or proteins showing altered levels in disease vs. control groups. This step lays the groundwork for downstream integration.
Step 2: Integration of Omics Layers
Next, data from different omics layers—such as transcriptomics and epigenomics—are combined. These integrations help correlate regulatory elements (e.g., methylation changes) with downstream effects like gene expression changes.
Step 3: Functional and Pathway Enrichment
Once significant molecular features are identified, they are analyzed to find biological pathways and functional categories that are overrepresented. This step helps contextualize the molecular alterations in terms of known cellular processes.
Step 4: Regulatory Element and Transcription Factor Mapping
Researchers then explore promoter and enhancer regions of genes to identify regulatory elements and the transcription factors (TFs) that might be controlling the expression of differentially regulated genes.
Step 5: Upstream Network Reconstruction
In this step, molecular networks are reconstructed to trace upstream regulators, such as signaling molecules or kinases, that may control the identified TFs. This reveals master regulators—key nodes that propagate signals across layers of cellular regulation.
Step 6: Drug Target and Candidate Prediction
Finally, researchers map these master regulators to existing drug databases or computational models to predict compounds that could modulate the identified regulatory network for therapeutic purposes.
While individual steps can be executed using a variety of publicly available and paid tools, integrating them into a coherent, reproducible workflow often requires custom scripting or multiple platforms.
Brief summary of the analysis performed by Genome Enhancer
The very first step (Step 1) of Genome Enhancer analysis aims to create a set of genes, which would describe the studied pathological process. In order to build this set, different actions are performed with different omics data types, depending on the initial input provided for the analysis.
After the gene set is identified, the Genome Enhancer pipeline proceeds to Step 2 to identify the regulatory regions (promoters and enhancers) of the selected genes.
On Step 3 the Genome Enhancer pipeline scans the regulatory regions, selected on Step 2, for transcription factors binding sites enrichment and identifies the modules of co-acting transcription factors using the TRANSFAC® database and the Match and Composite Module Analyst algorithms.
On Step 4 the Genome Enhancer pipeline performs network analysis and identifies the key nodes, responsible for the regulation of the studied pathological process. This is done by applying the Upstream Analysis approach towards the identified complexes of transcription factors with the use of TRANSPATH® database.
Based on the identified key nodes, the drug candidates search is performed on Step 5 of the Genome Enhancer pipeline with the use of HumanPSD™ database and PASS software.
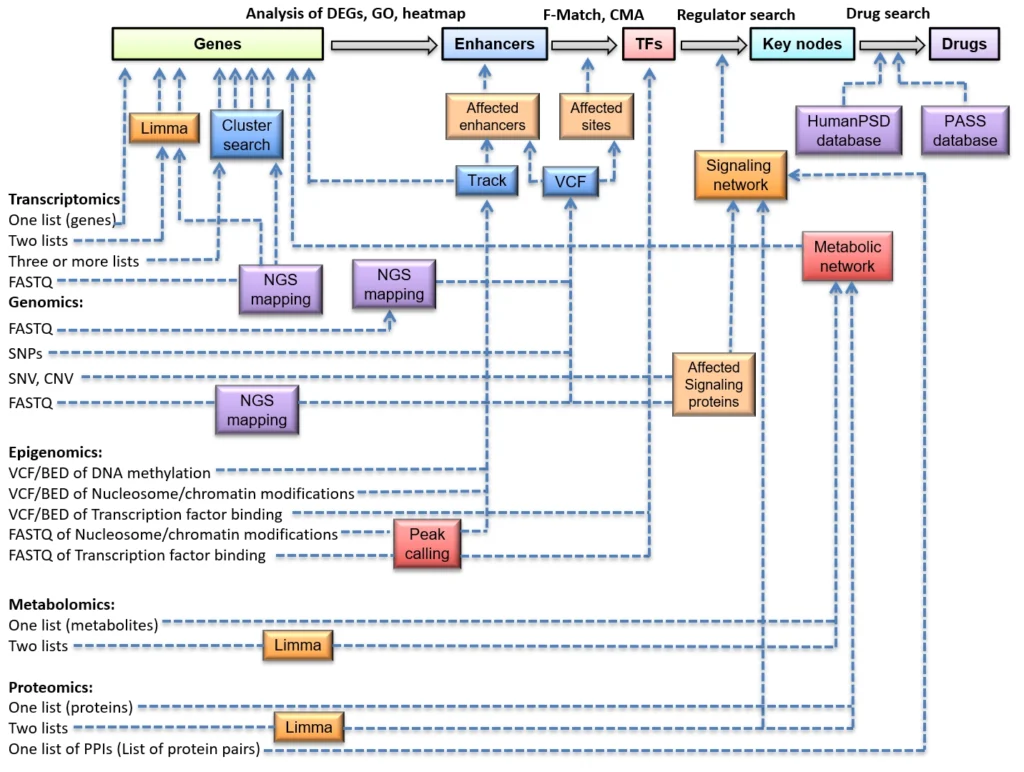
Detailed description of the performed analysis
Once the Genome Enhancer analysis was launched, the pipeline will create a workflow in respect to the analysis input. The workflow architecture depends on the number of selected conditions, which should be analyzed (the colored boxes from the data annotation diagram). For performing the analysis, Genome Enhancer will always take the latest versions of the databases, installed on the server.
Step 1. Constructing the gene set, which describes the studied pathological process
On this step the Genome Enhancer pipeline constructs the gene set, which describes the studied pathological process, from the input set of omics data.
Transcriptomics data
The transcriptomics data could have been loaded to Genome Enhancer in one of the following formats:
- Table data
- FASTQ files
- Affimetrix files
- Agilent files
- Illumina files
Table data
Genome Enhancer will separate the uploaded and submitted to the analysis transcriptomics table data into two types: (1) those files/columns, which came with numeric values (counts, logFC) and (2) those, which didn’t have numeric values and thus should be considered as a plain list of genes.
If transcriptomics data with numerical values was uploaded and submitted to the analysis, Genome Enhancer will check for which conditions (categories for comparison) this data is available.
If transcriptomics data with numerical values was submitted for only 1 condition, two options are possible:
- There were 1500 or less genes present in the input data
Then numerical values will be disregarded by Genome Enhancer and the provided data will be accepted as a plain gene list, which will be further taken to the next steps of analysis as the gene list, which describes the studied pathological process. - There were more than 1500 genes present in the input data
Then all columns in the table will be averaged and the following genes will be selected for further analysis:
top– 300 highly expressed genes
bottom– 300 low expressed genes
middle– 500 genes with medium expression, which will be considered as a background set (non-changed genes) in further analysis
In several further steps of analysis the following comparisons of expression of the genes (or other numerical values associated with the genes) will be made:top vs. middleandbottom vs. middle.
If transcriptomics data with numerical values was submitted for2 conditions, then the values from the 2 conditions will be compared to each other and the upregulated, downregulated and non-changed gene lists will be constructed.
If EdgeR method will be applied in the further analysis (see bellow), no additional normalization of transcriptomics data will be performed, otherwise the table will be normalized by performing Quantile normalization.
If data appeared to come in a non-logarithmic scale and no counts were present, the data will be log2 transformed using Transform Table method.
Then all data from all conditions will be integrated into one table. All empty cells of the resulting table will be filled in with average values throughout the lines using Table imputation.
The upregulated, downregulated and non-changed gene lists will be constructed according to the following rules:
If numerical transcriptomics data were present in two conditions:
- If each category contains at least 2 numerical columns, and not all of them are counts (integer numbers) then the Limma algorithm will be performed to compare the numerical values (e.g. gene expression values) between two conditions.
- If each of the compared conditions (categories for comparison) has 2 or more columns with counts data, the EdgeR algorithm will be performed to compare the numerical values (e.g. gene expression values) between two conditions.
- Otherwise, the Fold Change algorithm will be applied to the input numerical transcriptomics data.
If Limma or EdgeR were applied to the input data, then significant upregulated and significant downregulated genes will be selected for further analysis, as well as non-significant genes (background set). If the number of significantly changed genes will be over 300, then the top 300 significant genes will be taken for further analysis. If the number of significantly changed genes will be between 10 and 300, then all of these up or down regulated genes will be taken for further analysis. If the number of significantly changed genes will be less than 10, and the number of up or down regulated genes will be more than 300, then the top 300 up or down regulated genes will be taken for further analysis. If the number of significantly changed genes will be less than 10, and the number of up or down regulated genes will be less than 300, then all of these up or down regulated genes will be taken for further analysis. If the gene list will contain more than 2500 genes, the middle 500 genes will be considered as a background set. Otherwise, Genome Enhancer will take middle 20% of genes as a background set.
If Fold Change was applied to the input data, then upregulated and downregulated genes will be selected for further analysis, as well as non-changed genes (background set) in the same quantities as for Limma or EdgeR, described above.
In any case, after performing EdgeR, Limma or Fold Change, Genome Enhancer requires at least 10 genes with logarithm of the fold change values below zero and at least 10 genes with the logarithm of the fold change above zero to be present in the resulting data, otherwise further analysis of this data will not be performed.
Having constructed the lists of (significantly) up and down regulated genes, Genome Enhancer will proceed to the next step of the pipeline, where (significant) upregulated genes will be compared to the non-changed genes and (significant) downregulated genes will be compared to the non-changed genes. These two comparisons will form two independent branches of Genome Enhancer workflow, each of which will proceed to the next steps of the pipeline (find regulatory regions analysis (Step 2) and Match and CMA analysis (Step 3)).
If transcriptomics data with numerical values was submitted for 3 conditions (categories for comparison) selected during the analysis launch, two variants are possible:
- baseline was selected
- clustering was selected
In the (1) case Genome Enhancer will take all non-baseline conditions and will compare them to the baseline in the same way the analysis for 2 conditions was performed above. The results of all such comparisons will be tables with genes and, if applicable, corresponding numerical expression values. Each comparison will form an independent branch of Genome Enhancer workflow, which will proceed to the next steps of the pipeline (find regulatory regions analysis (Step 2) and Match and CMA analysis (Step 3)).
In the (2) case the cluster analysis will be launched.
First, the Variance filter analysis will be applied, which will go through all columns of the united transcriptomics table and identify genes with high and low variability in expression. The unchanged cluster will be constructed from the genes with low variance in expression and high variance cluster will be built from the genes with high variance in expression. Top 3000 genes will be taken from the cluster with high variance and 500 genes will be taken from the unchanged cluster for further analysis.
Then CRC analysis will be performed on the selected set of 3000 genes with high variability. The CRC analysis will generate as many clusters as it will identify.
On the results of CRC analysis (multiple clusters received) the three best clusters will be selected by launching of the CR cluster selector method, which will select the top 3 clusters of genes, not more than 300 genes and not less than 10 genes in each. Each of the three clusters will be then compared to the unchanged genes (500 genes with minimal variance), which will be treated as a control set. Each of these three comparisons will form an independent branch of Genome Enhancer workflow which will proceed to the next steps of the pipeline (find regulatory regions analysis (Step 2) and Match and CMA analysis (Step 3)).
If non-numeric transcriptomics data are loaded and submitted to the analysis, Genome Enhancer will compare the corresponding lists of genes of each of the conditions (categories for comparison) with the gene list in the baseline category and will select those genes, which do not belong to the genes of the baseline category for further steps of the pipeline (find regulatory regions analysis (Step 2) and Match and CMA analysis (Step 3)).
FASTQ data
If RNA-seq files in fastq format were uploaded and submitted to the analysis, Genome Enhancer will process them using the HISATalgorithm for alignment. After HISAT, HtSeq analysis will be applied to these files and a summary table with identified read counts will be generated if fastq files were present in two conditions (categories for comparison), with not less than 2 fastq files in each of the conditions. Afterwards this table will be processed with EdgeR method. Otherwise, if only one fastq file is used in the compared categories the Cufflinks analysis will be performed.
The resulting table with logarithms of fold changes and the computed p-values will be then treated as transcriptomics table data (described above).
Affymetrix, Illumina and Agilent data
If Affymetrix data was loaded (.CEL files), it will be normalized and converted into Ensembl genes. The initial file name and values will be kept in a newly created table together with the corresponding Ensembl and Affimetrix IDs. This action will be performed with all Affymetrix files that were uploaded and submitted for the analysis inside each of the conditions (categories for comparison). The resulting table will be treated as initial transcriptomics table data which is then analysed with Limma to detect the differentially expressed genes (described above).
Similar actions will be performed with Agilent and Illumina input data (Agilent normalization and Illumina normalization will be done).
Epigenomics data
If epigenomics data was loaded, Genome Enhancer will calculate the ChIP-seq peaks and will unite them in 1 track for each of the conditions (categories for comparison) which were specified during the analysis launch.
The epigenomics data could be loaded to Genome Enhancer in one of the following formats:
- Fastq files
- Bam files
- Tracks
- Tables with cg IDs and numerical values
If fastq file was uploaded and submitted for the analysis, Bowtie algorithm will be applied to it and respective bam file will be created. Next such files will be treated as if they were in bam format.
If bam file was loaded, the genome build of the file will be checked. Only hg38 bam files will be accepted by Genome Enhancer. For each of the valid bam files MACS 1.4 analysis will be performed, which will create the ChIP-seq peaks. All peaks inside one condition (category for comparison) will then be united. This will be done for all conditions (categories for comparison).
If track file was loaded, Genome Enhancer will check its genome build and will convert all non-hg38 tracks to hg38 genome build using the Liftover method. All tracks inside one condition (category for comparison) will be united. This will be done for all conditions (categories for comparison).
The performed actions will generate 1 resulting track, which will contain all respective ChIP-seq peaks, for each of the conditions (categories for comparison).
If no transcriptomics data was uploaded and submitted to the analysis and epigenomics data were present and submitted to the analysis, Genome Enhancer will take the unified track with peaks for each of the conditions and convert track to genes with Track to Gene set analysis, searching for peaks 1000bp downstream and 1000bp upstream of the gene. The resulting gene list is sorted by number of peaks intersected with the gene, then the top 300 genes will be considered as YES set and 500 bottom genes will be considered as NO set for each of the conditions. The pipeline will then proceed to the next steps of analysis (steps 2 and 3) with independent comparisons of YES set vs. NO set (top 300 genes vs. bottom 500 genes) for each of the conditions.
If table(s) with cg IDs (CpG loci IDs) and respective numerical values were loaded, their columns should be spread between the two conditions for comparison, representing the studied pathology and the control set. If several numerical columns were present, Fold Change will be calculated. Top 10 000 cg will be taken into further analysis and mapped to the track.
If the control set is not specified and only one condition under study contains columns with numerical data, values will be averaged among all selected columns and top 10000 cg will be taken into analysis and mapped to the track.
Tables with cg IDs (CpG loci IDs) without numerical data (CG lists) could also be processed. In this case CG lists are joined inside each category and then mapped to the tracks. If two categories with CG lists, experiment and control, are specified, control track will be subtracted from experiment track. If one, three or more categories with CG lists are specified, each track will be considered separately.
If no transcriptomics data was uploaded and table(s) with CpG loci IDs are present, Genome Enhancer will take the track(s) with methylation sites and convert track to genes with Track to Gene set analysis, searching for peaks 1000bp downstream and 1000bp upstream of the gene. The resulting gene list is sorted by number of methylation sites intersected with the gene, then the top 300 genes will be considered as considered as target genes of the studied pathology and compared to the unchanged genes (500 genes with minimal variance), which will be treated as a control set.
Proteomics data
If the gene set, which would describe the studied pathological process, was not constructed yet (from epigenomics or transcriptomics data because of their absence), Genome Enhancer will proceed to available proteomics data in order to generate such gene set on its basis.
Genome Enhancer will search throughout all conditions for proteomics data with numerical values (quantitative proteomics) and will perform the same actions on this data, as were described above for numerical transcriptomics data. All proteins will be converted to corresponding genes using the Convert table method.
The respective actions can be summarized as follows:
- If numerical proteomics data will be present only in one condition, Genome Enhancer will compare it to the housekeeping genes;
- If numerical proteomics data will be present in two conditions, they will be compared to each other;
- If numerical proteomics data will be present in three or more conditions, clustering or pairwise comparison will be performed depending on the clustering/baseline option selected during the analysis launch.
If the list of proteins (non-numeric proteomics) is present in the conditions, it will be converted to genes using the Convert tablemethod, then Genome Enhancer will compare the corresponding lists of genes of each of the conditions (categories for comparison) with the gene list in the baseline category and will select those genes, which do not belong to the genes of the baseline category for further steps of the pipeline.
If the gene set, which would describe the studied pathological process, was already constructed from epigenomics or transcriptomics data, than proteomics data will not be used for generating the initial list of genes.
On the next step of the pipeline the proteomics data will be used as so called “context set ” in the search for regulators in the signal transduction network. In the case if proteomics data are present only in one condition, the whole list of proteins is used as “context set”. If the proteomics with numeric values is present in two or more conditions, Fold change calculation is applied and proteins with the LogFC above zero are taken as the “context set”.
In the case of proteomics data without numerical values (simple list of protein IDs), if there is only one condition the whole protein list is used as the context set, but if there are two or more conditions then the proteins from the baseline condition will be disregarded and the proteins from the other conditions will be taken as the “context set”.
Genomics data
If the gene set, which would describe the studied pathological process, was not constructed yet (from epigenomics, transcriptomics or proteomics data because of their absence), Genome Enhancer will proceed to available genomics data in order to generate such gene set on its basis.
If a vcf files was loaded and submitted to the analysis, Genome Enhancer will check whether it comes in the hg38 genome build and, if not, then the input track will be converted to hg38 with the use of Liftover algorithm.
If an SNP list was loaded and submitted to the analysis, the SNP matching method will be applied and the respective VCF track will be retrieved as a result.
If a fastq file was loaded and submitted to the analysis, the respective vcf file will be generated using the workflow Find genome variants from full genome NGS.
All retrieved vcf tracks within one condition will be then joined.
If there were 2 conditions specified during the analysis launch, vcf track of Baseline will be subtracted from the vcf track of the other condition (studied pathology) using the Filter one track by another function. The Track to gene set method will be then applied to the resulting track and the set of genes from the mutation track will be retrieved.
Next the Mutations to genes with weights method will be applied.
The mutations present in the resulting track will be weighted depending on their localization:
- mutations, which got into the exon regions of genes, will receive the weight 0.7
- mutations, which got into the promoter regions of genes, will receive the weight 1.3
- all other mutations will receive the weight 1
The VCF track (Yes track) that was either provided as input or created by Genome Enhancer from SNP list or fastq files, is compared to Random VCF track (No track) of 10000 random human variations. On both tracks the score delta values are calculated (differences between PWM score values of the TF sites with the reference or with the alternative allele of the considered variation). For each variation we find then the maximal score delta values at each PWM leading either to the gain or to the loss of TF site (with the alternative allele). For selecting the maximum score delta values both directions of DNA strand are considered. Next, by going through all variations, two p-values for each PWM are computed: the p-value of site losses and the p-value of site gains. The p-values are computed using cumulative Binomial distribution estimating the random chances to observe the found high number of lost or gained TF sites in Yes track in the comparison to the No track. The PWM cut-offs are optimized to obtain the most extreme p-values. Top 20 best matrices by p-value from each: gained and lost sites are taken to further analysis. The mutation weights on the Yes track are calculated on the basis of these obtained 40 matrices. Each mutation is assigned with a respective matrix that got the maximum delta value either for the site gain or for the site loss (changed the binding affinity most significantly). This delta is then compared to other delta values that were computed for the respective matrix on the No track. The eventual weight that reflects the transcription factor binding affinity change caused by the mutation is calculated as follows:
w2 = -log10( NoGr / NoAll ), if NoGr > 0
w2 = -log10( 1.0 / ( 2.0 * NoAll ), if NoGr = 0
where NoGr is the number of deltas from the No track that appeared to be greater than the inspected delta and NoAll is the total number of deltas in the No track. The resulting track is then constructed that contains all sites of the initial Yes track together with the additional weights reflecting the transcription factor binding affinity change caused by the mutation.
The list of 40 matrices most affected by variations will be further used in composite modules search.
The genes, on which the mutations are localized, will also be weighted using the Calculate weighted Mutation Score analysis. This method will give genes the weights in respect to the number of mutations, which appeared to be localized on corresponding genes. In addition to that, genes belonging to TRANSPATH pathways will receive additional weight (a multiplier coefficient of 1.5 will be applied to all genes that happen to belong to TRANSPATH pathways). Also, this analysis will take into account the known gene-disease associations from the HumanPSD database for the diseases, which were specified during the analysis launch. In case no disease was specified during the analysis launch, the “Disease progression” pathology will be considered by default. Genes which happen to be associated with the diseases that were selected during the analysis launch will receive additional weight (a multiplier coefficient of 2 will be applied to genes that will be associated with the studied pathologies).
Total gene mutation weight is the sum of two weights: weight w1 of all variations located inside the gene body and in the gene flanking regions and weight w2 that reflects the transcription factor binding affinity change caused by the mutation. This weight is calculated by estimating the importance of a certain mutation in terms of gains or losses of binding sites caused by it.
The final list of mutations with their weights are used then for the analyses of TF binding sites at the later steps of the pipeline. In the case if no other type of data but only genomic data are present, the list of mutations is used then to generate the initial list of genes as follows.
The resulting table of genes and their weights will then be sorted by weights values and top 300 genes will be selected from for further analysis as top mutated genes. These genes will be saved to the table called ‘All mutated genes with description ranked’ which can be found in the results folder of the analysis from the Genome Enhancer Expert view. The selected 300 genes will then be filtered by genes in the baseline category (if two or more categories are used during the analysis launch) or taken as it is if only one category is used. This resulting list of genes will be compared to the list of housekeeping genes at the step of promoter analysis.
The mutation weights (w = w1+w2) will be also used to find the regulatory regions of the genes most affected by the variations/SNP. A sliding window of 1100 bp is used to scan through the intronic, 5’ and 3’ regions of the genes and a region is selected with the highest sum of the mutation weights.
Metabolomics data
If there was no transcriptomics data submitted to the analysis, the metabolomics data will be taken instead of it. If transcriptomic data was submitted to the analysis, the metabolomics data will be added to it.
If there were 2 conditions selected during the analysis launch, then only the list of metabolites from the non-baseline condition will be considered for further analysis.
If there were 1 or 3 and more conditions (and clustering option is used) selected during the analysis launch, then all lists of metabolites will be considered.
All tables of metabolites will be converted to Recon Substances using the Convert table function. Next, a list of genes encoding enzymes that are acting on these metabolites is obtained using the Match genes to metabolites function.
All genes will then be joined within one condition (category for comparison) and compared to the genes in baseline condition.
If there was only 1 numerical column present in the metabolomics data, Genome Enhancer will treat such data as a plain list of metabolites
If there were 2 numerical columns present in the metabolomics data, Genome Enhancer will apply Limma*, EdgeR* or Fold Change, in the same way as it was described for the transcriptomics data above.
*Limma and EdgeR will be calculated on Recon substances and only then the resulting up/down regulated metabolites will be converted to genes.
Then, similar to the way the transcriptomics data was treated, Genome Enhancer will find the significant upregulated, significant downregulated and unchanged genes if Limma or EdgeR were applied and upregulated and downregulated genes if Fold Change was applied.
The comparison will be further done between the (significantly) upregulated genes vs. housekeeping genes and (significantly) downregulated genes vs. housekeeping genes at the next steps of the pipeline.
The retrieved lists of genes from metabolomics analysis will be added to the previously received transcriptomics data, or, if no transcriptomics data were present, they will be taken as they are for further analysis.
Step 2. Identification of regulatory regions of the selected genes
On this step of Genome Enhancer pipeline the regulatory regions of the genes, which were selected on Step 1 for further analysis, will be identified. This will be done with Find regulatory regions method, which creates a track of regulatory regions for a set of input genes.
If epigenomics data was processed on the Step 1 of Genome Enhancer pipeline, then a united track with ChIP-seq peaks for each of the categories is constructed. This track will then be used in Find regulatory regions method for selecting the regulatory regions, which will tend to cover peaks intervals.
The Find regulatory regions method will select the peaks inside the input genes extended with flanks (shift on left gene bound position: -5000; shift on right gene bound position: 5000). Then intervals around the peak centers will be created, overlapped, and cut to input size (promoter from: -1000 and promoter to: 100, 1100bp in total). Finally, the closest to TSS interval will be selected as the result.
The TSS will be taken from Fantom/TSS database (CAGE TSS database: Fantom5-Tissue-hg38/TSS).
If no peaks are found around the gene, Fantom promoter will be used if tissue type was specified during the analysis launch and Ensembl promoter will be used if no tissue type was selected.
If genomics data was processed on the Step 1 of Genome Enhancer pipeline, then a track of identified mutations was constructed. In this case the Find regulatory regions with mutations method will be applied, which will create a track of regulatory regions for input genes using information about genomic variations from mutation track. The method will scan genes extended with flanking regions by window of input size: promoter from: -1000, promoter to: 100. It will find a position with highest sum of mutation weights that got into the scanned “window”. If no mutations will be found around the gene, Fantom promoter will be used if tissue type was specified during the analysis launch and Ensembl promoter will be used if no tissue type was selected.
If no epigenomics or genomics data was processed in the analysis, then the Find regulatory regions method will be simply performed on the list of genes, which describe the studied pathological process (list of genes, that was constructed on Step 1 of Genome Enhancer pipeline).
The retrieved regulatory regions will be used in the next step of Genome Enhancer pipeline for identification of transcription factors binding sites and their complexes.
Step 3. Identification of transcription factor binding sites and their complexes
On this step Genome Enhancer pipeline the regulatory regions, which were identified on Step 2, will be scanned for transcription factors binding sites enrichment and the modules of co-acting transcription factors that regulate the genes of the studied pathological process will be identified using the TRANSFAC® database and the Match and Composite Module Analyst (CMA) algorithms. In the Match and CMA algorithms the frequencies of TFBS and TFBS composite modules are compared in the regulatory regions of foreground set of genes (Yes-set) to the regulatory regions of the background set of genes (No-set). In each comparison of a category to the baseline two Yes-sets are formed – one is the list of up-regulated genes and second is the list of down-regulated genes. The No-set is prepared from the non-changed genes. In the case of one category only, the Yes-sets are formed from thetopandbottomgenes, and No-set is formed from the middle genes (in the case of numerical values associated with the genes). In the cases of analysis of gene lists without numerical values (see above) the No-set is formed from the housekeeping genes.
In details, the Site search on track method is applied to the regulatory regions, selected on step 2 of the Genome Enhancer pipeline, to predict the transcription factor binding sites. The retrieved result is further optimized with the use of Site search result optimizationmethod, which tunes the matrix weights to minimize p-values.
Independently, in the case of genomic data are used, the revealed mutations that are mapped to the regulatory regions of the differentially expressed genes are analyzed by the method Compare TFBS mutations, which identified TFBS that are lost or gained due to the mutations. The list of PWMs of the top significant lost or gained TFBS is used then in CMA analysis for specifying the search of the composite modules.
Next, the CMA (Composite Module Analyst) analysis is be performed by running the Construct composite modules on tracks method. Composite modules are combinations of binding sites common for promoters of functionally related genes and responsible for the major component of the gene expression pattern of these genes. If genomics data was present as input, the matrices which were identified as matrices causing the significant change in the transcription factor binding affinity as the result of the observed mutation, will be taken into account. Each of the resulting CMA composite modules will have to include at least one such matrix. The retrieved modules will be used on the next step of the Genome Enhancer pipeline for identification of the key nodes, responsible for the regulation of the studied pathological process.
Further information on methods for identification of transcription factor binding sites and their complexes can be found in the ‘Methods for the analysis of enriched transcription factor binding sites and composite modules’ subsection of the Methods section in Genome Enhancer analysis report.
Step 4. Identification of key nodes, responsible for the regulation of the studied pathological process
On this step of Genome Enhancer pipeline the network analysis is performed and the key nodes, which are responsible for regulation of the studied pathological process, are identified.
For this the Molecular networks regulator search method will be applied to the set of transcription factors, selected on the Step 5 of Genome Enhancer pipeline. The regulator search will be performed with the use of TRANSPATH® database for molecules upstream of the input list of transcription factors. As the result, this method will generate a set of proteins or their encoding genes, which were predicted to play a key role in regulating a maximal number of transcription factors from the input list.
If proteomics data was processed on Step 1 of the Genome Enhancer pipeline, the list of context proteins, that refer to the studied pathological process, will be used for weighting the respective molecules on the reconstructed network of intracellular reactions. Proteins, belonging to the list of proteins, which characterize the studied pathological process, will have higher priority for appearing on the reconstructed signaling network.
Opposite to that, a list of heavily mutated signaling proteins, that were extracted from genomics data in case of its presence on Step 1 of the Genome Enhancer pipeline, will be excluded from the constructed signaling network due to the loss of their function in the studied pathological process.
Further information on methods for identification of key nodes, responsible for regulation of the studied pathological process, can be found in the ‘Methods for finding master regulators in networks’ subsection of the Methods section in Genome Enhancer analysis report.
Step 5. Identification of prospective drug candidates
Based on the key nodes, identified on Step 4 of Genome Enhancer pipeline, the drug candidates search will be performed onStep 5of the pipeline with the use of HumanPSD™ database and PASS software.
For identification of already approved drugs and drugs undergoing clinical trials for the studied pathology and for other diseases the PSD pharmaceutical compounds analysis will be launched by Genome Enhancer pipeline. This method will seek for the optimal combination of molecular targets among the set of the input genes (key nodes, identified on Step 4 of the Genome Enhancer pipeline), that can potentially interact with pharmaceutical compounds from a library of known drugs, using information from HumanPSD™database. As a result, this method will provide the identified lists of prospective drug targets and respective treatments that were proven in clinical trials to affect the identified key nodes and thus potentially block the studied pathological process.
Then the chemoinformatics analysis will be applied with the use of PASS software and chemoinformatically predicted prospective drug targets and treatments will be identified by Genome Enhancer pipeline. This will be done by performing the Pharmaceutical Compounds analysis which will seek for the optimal combination of molecular targets among input genes (key nodes, identified on Step 4 of the Genome Enhancer pipeline), that potentially interact with pharmaceutical compounds from a library of known drugs and biologically active chemical compounds predicted with the cheminformatics tool PASS. As a result, this method will provide the identified lists of prospective drug targets and respective treatments that were chemoinformatically predicted to affect the identified key nodes and thus potentially block the studied pathological process.
Further information on methods for identification of prospective drug candidates can be found in the ‘Methods for analysis of pharmaceutical compounds’ subsection of the Methods section in Genome Enhancer analysis report.
For further assistance please contact [email protected]
Acceptable input data formats
Genome Enhancer works with genomics, transcriptomics, epigenomics, proteomics and metabolomics input data types of the following formats:
Transcriptomics (RNA-seq, microarrays)
*.txt, *.csv, *.xls (table with gene identifiers)
*.CEL (affymetrix)
*.txt (special agilent format)
*.txt (special illumina format)
*.fastq
Epigenomics (ChIP-seq)
*.fastq
*.bam (hg38 only)
*.bed (hg38 only)
*.txt (table with illumina methylation probe ids, cg*)
Genomics
*.vcf
*.txt, *.csv, *.xls (table data with SNP identifiers, rs*), *.tsv
*.fastq
Proteomics
*.txt, *.csv, *.xls (table with protein identifiers)
Metabolomics
*.txt, *.csv, *.xls (table with the list of metabolites from chebi database, e.g. CHEBI:57316)
Detailed Report
Detailed Report with Everything You Need for Your Paper
Transparent, Traceable, and Publication-Ready Results — All in One Document
Genome Enhancer doesn’t just deliver final outcomes—it provides a comprehensive, scientifically structured report that includes everything you need to publish your results or present them to collaborators. From raw omics input to prioritized drug targets, every analytical step is clearly documented with methods, references, diagrams, and traceable outputs.
Whether you’re preparing a manuscript, funding proposal, or internal review, this report gives you a full breakdown of the analytical logic and results, including intermediate stages of analysis that can support supplementary materials or reviewer questions.
No more black-box outputs or vague summaries. You get clarity, transparency, and rigor.
What’s Inside the Genome Enhancer Report
Full Methodology and References
Each step of the pipeline is described in detail, from promoter analysis to master regulator detection. The report includes:
- Explanation of algorithms (e.g., MATCH™, CMA, PASS)
- Database sources (e.g., TRANSFAC®, TRANSPATH®, HumanPSD™)
- Proper scientific references for all tools and resources used
Final Results + Intermediate Steps
In addition to high-level conclusions, the report walks you through:
- Differential expression or methylation data
- Detected transcription factors
- Pathway reconstruction steps
- Master regulator identification
- Drug prioritization criteria and scores
Direct Access to Full Data
The report includes:
- Hyperlinked tables with full gene lists, pathways, TFs, and drug predictions
- Clickable diagrams for molecular networks and regulatory trees
- Export options for figures and data in formats suitable for publication
Visualizations Ready for Your Manuscript
High-quality figures such as:
- Master regulator signaling networks
- Transcription factor enrichment maps
- Pathway overlays with affected genes
- Drug-target interaction tables with supporting evidence
Why It Matters
- Supports scientific reproducibility
- Saves weeks of manuscript preparation time
- Reduces back-and-forth with collaborators or reviewers
- Gives you confidence in every step of the result chain
- Provides a traceable audit trail for grant proposals, thesis work, or clinical reporting
Example Output: Automatically Generated Report PDF
Get free reports and case studies to your Email
Benefits of the One-Click Workflow
- Shortens analysis time from weeks to hours
- Removes the need for custom scripting or pipeline assembly
- Enables non-bioinformaticians to analyze complex datasets
- Produces standardized, publication-ready results
- Supports clinical translation with prioritized treatment suggestions
- Identifies activated targets in the examined patient data and suggests known and re-purposing drugs.
Genome Enhancer workflow execution
Genome Enhancer uses only three steps to launch the analysis
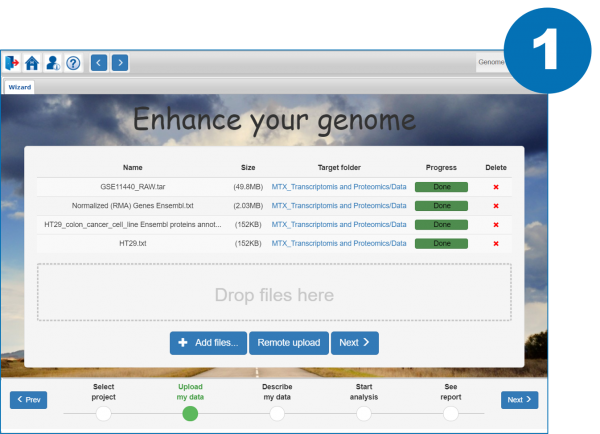
1. Upload your data to the server and specify the import options (data type)
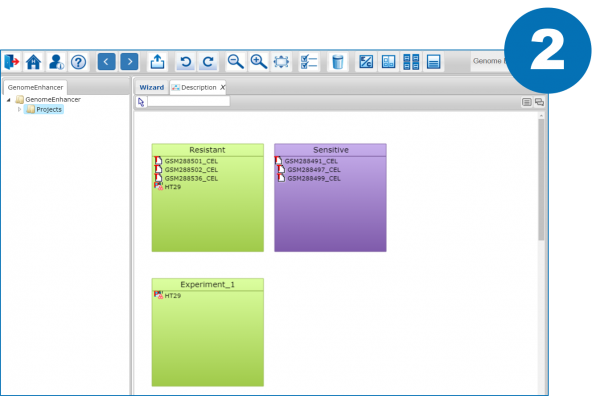
2. Split your data by the conditions you want to compare
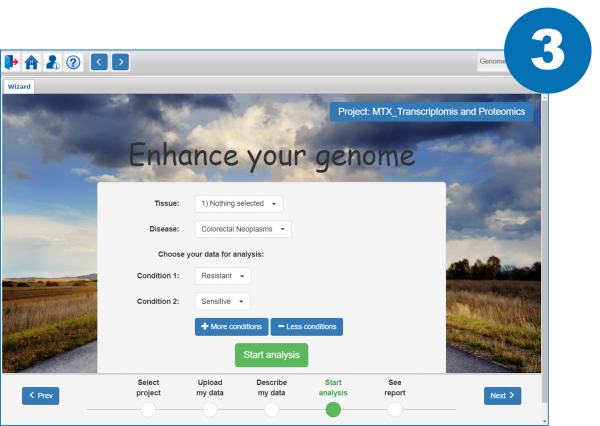
3. Launch the analysis by specifying the conditions to be compared and the disease and tissue types (optional)
The analysis report will be ready shortly. Depending on your input data, it will include lists of differentially expressed or mutated genes; transcription factors, regulating those genes; reconstructed signaling network of the studied pathological process; potential drug targets and corresponding known drugs and re-purposing drugs, which may be effective in the studied case, as well as further cheminformatically predicted drug-like compounds. The report also contains description of analysis methods used and the references.
Acceptable input data formats
Genome Enhancer works with genomics, transcriptomics, epigenomics, proteomics and metabolomics input data types of the following formats:
Transcriptomics (RNA-seq, microarrays)
*.txt, *.csv, *.xls (table with gene identifiers)
*.CEL (affymetrix)
*.txt (special agilent format)
*.txt (special illumina format)
*.fastq
Epigenomics (ChIP-seq)
*.fastq
*.bam (hg38 only)
*.bed (hg38 only)
*.txt (table with illumina methylation probe ids, cg*)
Genomics
*.vcf
*.txt, *.csv, *.xls (table data with SNP identifiers, rs*), *.tsv
*.fastq
Proteomics
*.txt, *.csv, *.xls (table with protein identifiers)
Metabolomics
*.txt, *.csv, *.xls (table with the list of metabolites from chebi database, e.g. CHEBI:57316)
Files of one data format can be uploaded in a .zip archive
Drug repurposing
Insights from Multi-Omics and Machine Learning
Drug repurposing offers a high-impact opportunity to identify new therapeutic uses for existing drugs—especially in complex, treatment-resistant cancers. Genome Enhancer is a machine learning–powered platform that applies upstream regulatory modeling to multi-omics data, integrating RNA-seq and DNA methylation to unravel hidden mechanisms behind disease phenotypes.
In a case study on cisplatin-resistant ovarian cancer, Genome Enhancer was used to analyze transcriptomic and epigenomic profiles of treated and untreated cancer cells. The goal was: to identify druggable targets and repurposing-ready drugs that could help overcome resistance.
The analysis revealed several transcription factors (EP300, NFYA, RELA, RXRA, HSF1) driving gene expression changes in the resistant phenotype. Through regulatory network reconstruction, key master regulators upstream of these TFs were identified—including PDGFRA, VRK1, and the CDK1–Cyclin B1 complex.
These master regulators represent pivotal control points in the disease-specific signaling network—and offer promising entry points for therapeutic intervention.
How Master Regulators Are Identified
- RNA-seq, ChIP-seq, ATAC-seq, DNA-methylation and other multi-omics data Input:
The workflow begins with identification of differentially expressed and epigenetically regulated genes from patient-derived omics data. - TF Binding Site Prediction:
Promoter and enhancer regions of these genes are analyzed using the TRANSFAC® database and MATCH™/CMA algorithms to detect overrepresented transcription factor binding sites (TFBSs) and their combinations. - Pathway Tracing:
Using TRANSPATH®, the tool reconstructs upstream signaling pathways that could explain the activation of these TFs. - Master Regulator Detection:
Specialized graph algorithms scan the pathway network to pinpoint nodes at the top of these cascades—called master regulators. These are molecules whose activity explains the global shift in transcriptional programs seen in the pathology.
How Drugs Are Prioritized
- Druggability Assessment:
Each identified master regulator is checked against the HumanPSD™ database for known drug–target relationships and literature evidence. - Biological Activity Prediction (PASS):
For regulators with no direct drug hits, the system uses PASS (Prediction of Activity Spectra for Substances) to identify chemical compounds likely to interact with those targets based on structural similarity and known activity profiles. - Compound Filtering:
The final drug list includes:
- Approved FDA drugs with known efficacy in related pathways
- Investigational compounds with high predicted interaction scores
- Prioritization based on network-level modulation and existing pharmacokinetics/safety data
- Approved FDA drugs with known efficacy in related pathways
In this study, Paclitaxel and Etoposide were highlighted as promising candidates for repurposing in cisplatin-resistant ovarian cancer based on their predicted ability to modulate the disease-specific regulatory network.
What You’ll Gain
- A full systems-level view of molecular dysregulation in cisplatin resistance
- A prioritized list of master regulators and actionable targets
- Repurposing candidates from FDA-approved and investigational drug libraries
- A replicable methodology for >3,900 diseases using Genome Enhancer
Master Regulator Map Example (Generated Automatically)
Below we represent schematically the main mechanism of the studied pathology. The pipeline proposed the following key master regulators:
- PDGFRalpha
- vrk1
- Cdk1-isoform1(h):cyclinB1-isoform1
This result allows us to suggest the following schema of affecting the molecular mechanism of the studied pathology:
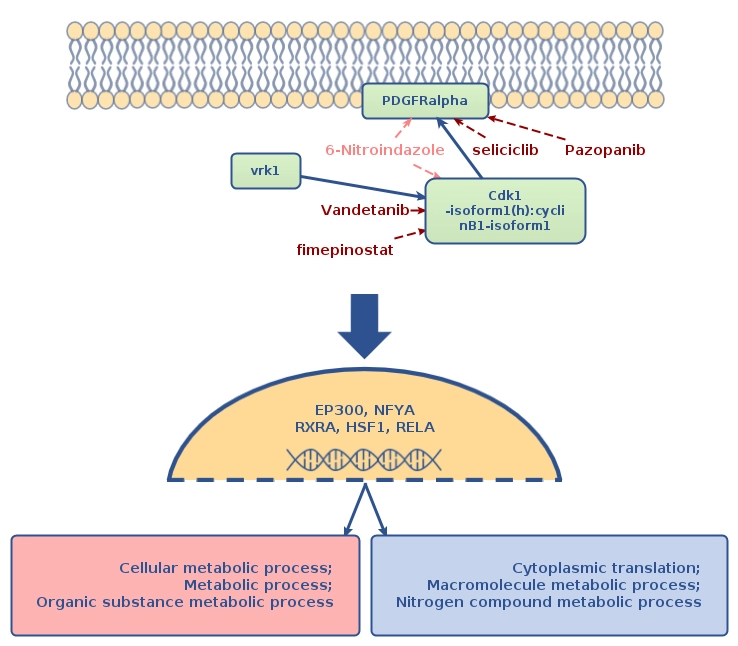
Drugs which are shown on this schema: Vandetanib, 6-Nitroindazole, seliciclib, fimepinostat and Pazopanib, should be considered as a prospective research initiative for further drug repurposing and drug development. These drugs were selected as top matching treatments to the most prospective drug targets of the studied pathology, however, these results should be considered with special caution and are to be used for research purposes only, as there is not enough clinical information for adapting these results towards immediate treatment of patients.
The drugs given in dark red color on the schema are FDA approved drugs or drugs which have gone through various phases of clinical trials as active treatments against the selected targets.
The drugs given in pink color on the schema are drugs, which were cheminformatically predicted to be active against the selected targets.
What You’ll Gain
- A reproducible systems biology workflow in a single click
- Integration of regulatory genomics and pathway modeling
- High-confidence drug target and compound predictions
- Visual and interpretable output for biological teams and decision makers
Report examples
You can view various analysis report examples generated by Genome Enhancer on the basis of different omics input data types and various origins of the studied pathologies:
- Colorectal Cancer (Personalized patient data) — Genomics, VCF
- MTB (Molecular Tumor Board) report example for colorectal cancer patient — Genomics, VCF
- Esophageal Squamous Cell Carcinoma (GSE32424) — Transcriptomics, FASTQ
- IFN-alpha induction (GSE31193) — Transcriptomics, LogFC Table
- Lung cancer, treatment by TGF (ST000010) — Metabolomics, Table
- Osteosarcoma, neoplasm metastasis (GSE66789) — Transcriptome + Proteome, RNA-seq + Mass-spec proteomics
- Ovarian cancer, cisplatin-resistance (GSE15709) — Transcriptomics + Epigenomics, CEL + BED
- SNP associated with Diabetes Mellitus — Genomics, SNP list
- Parkinson disease, induced a-Syn expression in SH-SY5Y cells (GSE145804) — Transcriptomics, LogFC Table
- Non-Small Cell Lung Carcinoma (NCI-H1975) — Genomics, VCF
- MTB (Molecular Tumor Board) report example for non-small cell lung carcinoma (NCI-H1975) — Genomics, VCF
- Hypertension (GSE157131) — Epigenomics, cg lists
Video
Below you will find a compilation of our videos referring to different aspects of Genome Enhancer.
This video introduces you to the fully automated pipeline for the easy bioinformatics analysis of multi-omics data. (2:02 min)
This video shows how Genome Enhancer results can be interpreted for further use in clinic on the example of sensitivity prediction towards VEGFA-targeted therapy for three colorectal cancer patients. (5:51 min)
References
- Lloyd, Katie, et al. “Using systems medicine to identify a therapeutic agent with potential for repurposing in Inflammatory Bowel Disease.” Disease models & mechanisms (2020).
- Kel, Alexander, et al. “Walking pathways with positive feedback loops reveal DNA methylation biomarkers of colorectal cancer.” BMC bioinformatics 20.4 (2019): 119.
- Kolpakov, Fedor, et al. “BioUML: an integrated environment for systems biology and collaborative analysis of biomedical data.” Nucleic acids research 47.W1 (2019): W225-W233.
- Boyarskikh, Ulyana, et al. “Computational master-regulator search reveals mTOR and PI3K pathways responsible for low sensitivity of NCI-H292 and A427 lung cancer cell lines to cytotoxic action of p53 activator Nutlin-3.” BMC medical genomics 11.1 (2018): 12.
- Boyarskikh, U. A., et al. “Master-regulators driving resistance of non-small cell lung cancer cells to p53 reactivator Nutlin-3.” Virtual Biology 4 (2017): 1-31.
How to cite Genome Enhancer
Kel, A., Boyarskikh, U., Stegmaier, P., Leskov, L.S., Sokolov, A.V., Yevshin, I., Mandrik, N., Stelmashenko, D., Koschmann, J., Kel-Margoulis, O. and Krull, M. Walking pathways with positive feedback loops reveal DNA methylation biomarkers of colorectal cancer. BMC bioinformatics. Cambridge (UK): RSC Publishing. 2019;20(Suppl 4):119:1-20. Link
Disclaimer
The results of Genome Enhancer analysis, contained in any of the reports produced by this pipeline, are intended for research use only and should not be used for medical or professional advice. GeneXplain GmbH makes no guarantee of the comprehensiveness, reliability or accuracy of the information contained in the reports generated by Genome Enhancer.
Decisions regarding care and treatment of patients should be fully made by attending doctors. The predicted chemical compounds listed in the reports are given only for doctor’s consideration and they cannot be treated as prescribed medication. It is the physician’s responsibility to independently decide whether any, none or all of the predicted compounds can be used solely or in combination for patient treatment purposes, taking into account all applicable information regarding FDA prescribing recommendations for any therapeutic and the patient’s condition, including, but not limited to, the patient’s and family’s medical history, physical examinations, information from various diagnostic tests, and patient preferences in accordance with the current standard of care. Whether or not a particular patient will benefit from a selected therapy is based on many factors and can vary significantly.
The compounds predicted to be active against the identified drug targets in the reports are not guaranteed to be active against any particular patient’s condition. GeneXplain GmbH does not give any assurances or guarantees regarding the treatment information and conclusions given in the reports. There is no guarantee that any third party will provide a refund for any of the treatment decisions made based on these results. None of the listed compounds was checked by Genome Enhancer for adverse side-effects or even toxic effects.
The analysis reports contain information about chemical drug compounds, clinical trials and disease biomarkers retrieved from the HumanPSD™ database of gene-disease assignments maintained and exclusively distributed worldwide by geneXplain GmbH. The information contained in this database is collected from scientific literature and public clinical trials resources. It is updated to the best of geneXplain’s knowledge however we do not guarantee completeness and reliability of this information leaving the final checkup and consideration of the predicted therapies to the medical doctor. In all cases, the end user (including researchers and medical doctors) accepts full responsibility for all risks associated with using of information, contained in the reports generated by Genome Enhancer.
The scientific analysis underlying the Genome Enhancer reports employs a complex analysis pipeline which uses geneXplain’s proprietary Upstream Analysis approach, integrated with TRANSFAC® and TRANSPATH® databases maintained and exclusively distributed worldwide by geneXplain GmbH. The pipeline and the databases are updated to the best of geneXplain’s knowledge and belief, however, geneXplain GmbH shall not give a warranty as to the characteristics or to the content and any of the results produced by Genome Enhancer. Moreover, any warranty concerning the completeness, up-to-dateness, correctness and usability of Genome Enhancer information and results produced by it, shall be excluded.
The results produced by Genome Enhancer, including the analysis reports, severely depend on the quality of input data used for the analysis. It is the responsibility of Genome Enhancer users to check the input data quality and parameters used for running the Genome Enhancer pipeline.
Note that the text given in the reports is not unique and can be fully or partially repeated in other Genome Enhancer analysis reports, including reports of other users. This should be considered when publishing any results or excerpts from the reports. This restriction refers only to the general description of analysis methods used for generating the reports. All data and graphics referring to the concrete set of input data, including lists of mutated genes, differentially expressed genes/proteins/metabolites, functional classifications, identified transcription factors and master regulators, constructed molecular networks, lists of chemical compounds and reconstructed model of molecular mechanisms of the studied pathology are unique in respect to the used input data set and Genome Enhancer pipeline parameters used for the current run.
