TRANSFAC DISEASES
Identify drug targets and disease biomarkers
TRANSFAC® DISEASES is an integrated solution for exploring the molecular basis of human diseases and identifying clinically relevant targets. It combines curated regulatory and pathway knowledge with advanced analysis capabilities to support translational research.
The package includes the content of TRANSFAC® BASIC and TRANSFAC® PATHWAYS, and is further extended with the Human Proteome Survey Database (HumanPSD) and Genome Enhancer.
HumanPSD is a comprehensive catalog of human proteins, their complexes, and orthologs in mouse and rat. Its primary focus is on the association of human proteins with diseases, as well as their potential application as biomarkers and therapeutic targets.
Genome Enhancer is a fully automated pipeline for patient omics data analysis. It reconstructs disease-specific molecular mechanisms to identify master regulators, potential drug targets, and candidate therapeutic strategies.
TRANSFAC® DISEASES enables researchers to fully leverage integrated gene regulation and disease data within a unified framework, supporting a deeper understanding of disease mechanisms and facilitating data-driven target discovery.
HumanPSD
The Human Proteome Survey Database (HumanPSDTM) is a catalog of proteins and their complexes from human cells, plus their orthologs from mouse and rat sources.
Its main focus is on the association of human proteins with diseases as well as on their potential use as biomarkers.
Drugs targeting human proteins are reported. In addition, information can be retrieved on the molecular functions, biological roles, localization, and modifications of proteins, expression patterns across cells, tissues, organs, and tumors, consequences of gene mutations in mice, and the physical and regulatory interactions between proteins and genes. HumanPSDTM is available for online access fully integrated with TRANSPATH®.
Picture of HumanPSD
Introduction
Description
Synonyms
Biomarker Associations
Diseases associated with MYC
Inherit MYC mutations
Drug Interactions
Drug(s) targeting MYC
Gene Ontology
Molecular function
Biological process
Cellular component
Expression
Tissue expression
Regulation of MYC expression
Mutant Phenotype
Mutant phenotype of closely related homolog(s)
Pathways & interactions
Pathways
Protein-protein interactions
Events acting on MYC
Events triggered by MYC
Transcriptional Regulation
Add a subscription to TRANSFAC® and this report will display additional information
RNA Features
Overview of RNA sequence
Protein Features
Overview of protein sequence and structure
Post-translational modifications of MYC protein
View complex containing MYC protein
Identifiers
Accessions mapped to this record
Annotations
Description
Editor’s Notes
Disease related
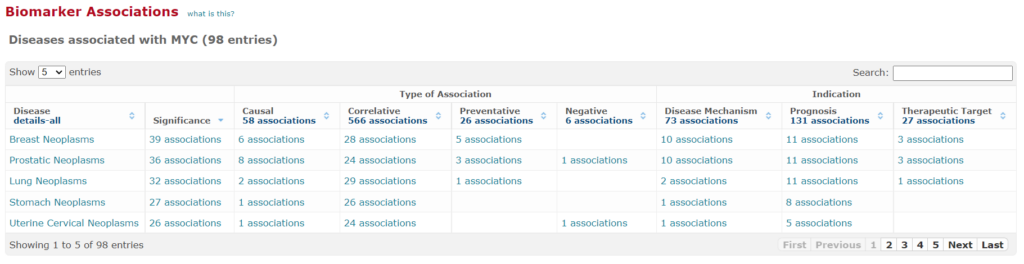
Biomarker / disease associations
A tabular summary of literature-derived relationships between human genes and gene products with human diseases is given.
These associations are clearly sorted according to their type, e.g. whether a gene/protein has a causal relationship with a disease to develop, or whether it is merely correlative, etc.
Drugs and targets
HumanPSD reports over 55,000 drug targets and associated with them over 10,000 drugs.
Disease mechanism
HumanPSD reports description of disease molecular mechanisms for over 3,900 human diseases.
HumanPSD also reports inferred disease-disease relationships on the basis of shared Causal and Preventative biomarker genes. Disease networks are derived from these relationships and organized into clusters with apparent biomedical relevance. The inferred disease-disease associations coincided with known clinical correlations such as comorbidities or known disease etiologies.
Adenocarcinoma
Disease similarity map:
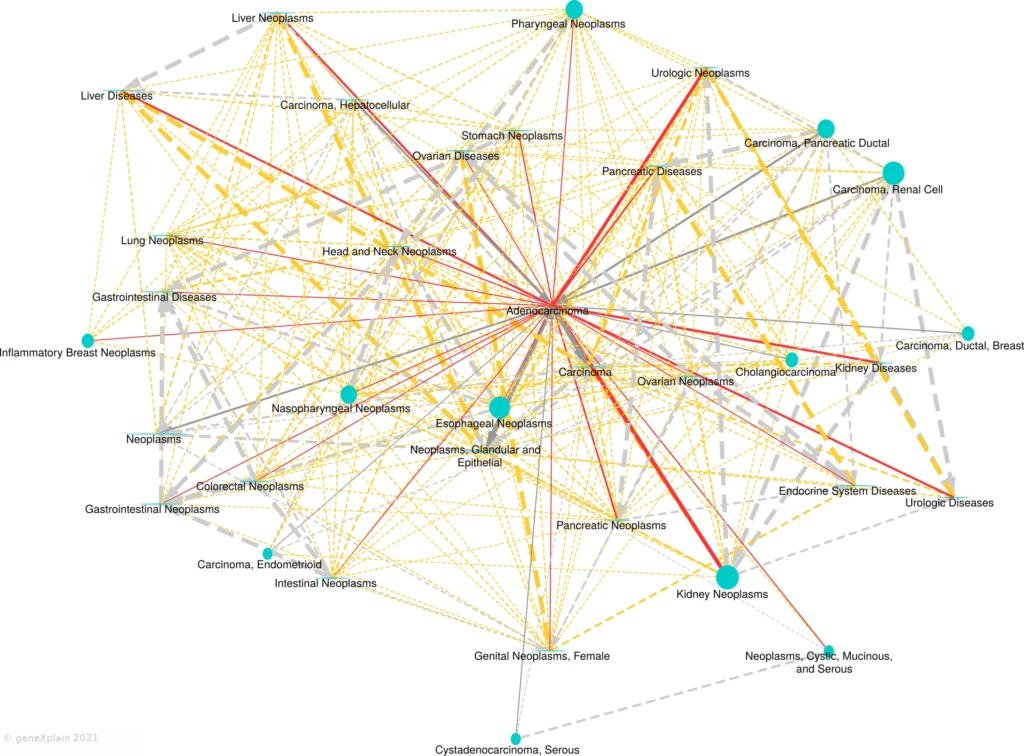
Legend:
This Disease Similarity Map connects diseases (nodes) with edges on the basis of common causal biomarker genes (edges are shown for FDR < 0.05 and overlap size >= 2). The primary disease is represented by a red diamond shape, neighboring diseases by blue circles. Solid edges connect the primary disease to neighboring diseases. Dashed edges connect neighboring diseases. Undirected solid red edges connect the primary disease in the center with similar diseases (neighbors), undirected dashed orange edges connect similar neighbors. Gray arrows point from a child disease to the parent disease according to the MeSH hierarchy. Edge widths are proportional to the statistical significance of association and node sizes are proportional to the number of causal biomarker genes of the disease.
Clinical trials
HumanPSD reports over 1,100,00 clinical trial – disease connections extracted from ClinicalTrials.gov and AACT databases, and also from the registries and data partners contributions to the OpenTrials project.
Clinical trials for Lung Neoplasm
Here is a screenshot of the information about clinical trials for Lung Neoplasms:
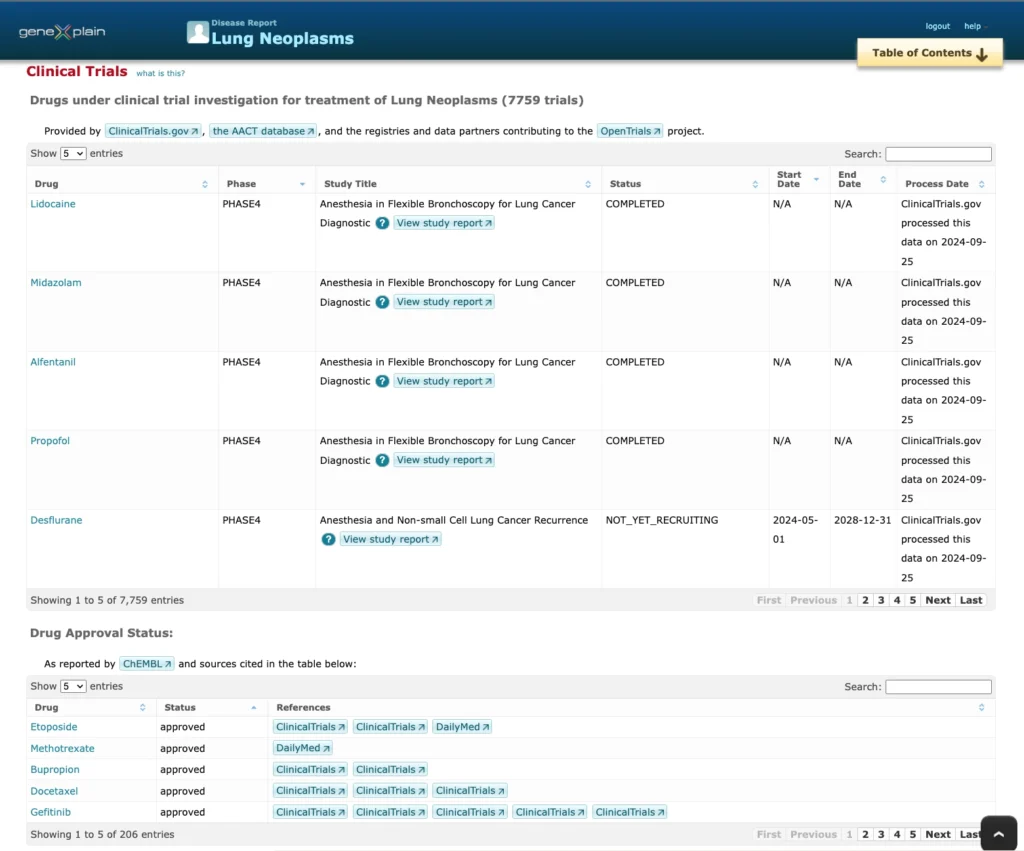
Key features
Benefits
Videos
Drug report and pathway report in HumanPSD+TRANSPATH® – this video demonstrates how drug report looks like in the online interface of HumanPSD+TRANSPATH® database. Target molecules, corresponding to a particular drug, are demonstrated, and pathways, in which these molecules are involved, are shown in a very detailed way, with all the underlying reactions, from which the corresponding pathways are constructed. Clinical trials info is also shown for each particular drug and for certain cases even metabolizing enzymes and associated pathways info is also available.
Disease report overview in HumanPSD+TRANSPATH® – this video demonstrates how disease report looks like in the online interface of HumanPSD+TRANSPATH® database. Disease biomarkers are shown (genes or proteins or miRNAs) with the type of association and type of indication info. All respective references are demonstrated. Disease similarity maps are also shown (these maps are based on the similarity of different diseases in respect to the common biomarkers that they share).
Disease similarity maps by common biomarkers in HumanPSD+TRANSPATH® – this video shows the overview of disease similarity maps. Common biomarkers of different diseases are displayed on these schemas. The searched disease is placed in the center of the map. The maps have the following color code: diseases sharing common biomarkers are connected with red lines; diseases that have ontological relationships according to the MeSH ontology are connected by gray lines.
Browsing pathways in HumanPSDTM+TRANSPATH®
Locus report in HumanPSDTM+TRANSPATH®
Recent applications
Selection of articles reporting about HumanPSD applications:
- Kawashima Y., Nagai H., Konno R., Ishikawa M., Nakajima D., Sato H., Nakamura R., Furuyashiki T., Ohara O. (2022) Single-Shot 10K Proteome Approach: Over 10,000 Protein Identifications by Data-Independent Acquisition-Based Single-Shot Proteomics with Ion Mobility Spectrometry. J Proteome Res. 21(6), 1418–1427. Link
- Lim, J. S., Ibaseta, A., Fischer, M. M., Cancilla, B., O’Young, G., Cristea, S., Luca, V. C., Yang, D., Jahchan, N. S., Hamard, C., Antoine, M., Wislez, M., Kong, C., Cain, J., Liu, Y. W., Kapoun, A. M., Garcia, K. C., Hoey, T., Murriel, C. L., & Sage, J. (2017). Intratumoural heterogeneity generated by Notch signalling promotes small-cell lung cancer. Nature, 545(7654), 360–364. Link
- Reales‐Calderón, J. A., Aguilera‐Montilla, N., Corbí, Á. L., Molero, G., & Gil, C. (2014). Proteomic characterization of human proinflammatory M1 and anti‐inflammatory M2 macrophages and their response to Candida albicans. Proteomics, 14(12), 1503-1518. Link
- Martínez‐Solano, L., Nombela, C., Molero, G., & Gil, C. (2006). Differential protein expression of murine macrophages upon interaction with Candida albicans. Proteomics, 6(S1), S133-S144. Link
Publications
Selection of publications authored by the geneXplain team:
- Stelmashenko, D., Kel-Margoulis, O., Apalko, S., & Kel, A. (2022) GENE NETWORKS AND DRUGS. WHAT CAN WE LEARN USING BIO-AND CHEMOINFORMATICS? In XXXVIII Symposium of Bioinformatics and Computer-Aided Drug Discovery (pp. 21-21).
- Kalya, M., Kel, A., Leha, A., Altynbekova, K., Wingender, E., Beißbarth, T. (2022). Machine Learning based Survival Group Prediction in Glioblastoma. Preprints 2022, 2022020051. Link
How to cite HumanPSD
Michael H, Hogan J, Kel A, Kel-Margoulis O, Schacherer F, Voss N, Wingender E. Building a knowledge base for systems pathology. Brief Bioinform. 2008 Nov;9(6):518-31. Link
