TRANSFAC PATHWAYS
Reconstruct signal transduction network controlling your genes.
Understanding which signaling or metabolic pathways are activated under specific experimental conditions is a common objective in life science research. A related challenge is identifying the pathways responsible for driving an observed phenotype.
Both questions can be effectively addressed using the analytical tools available in the TRANSFAC PATHWAYS package, enabling researchers to connect experimental data with underlying biological mechanisms in a systematic and interpretable way.
The TRANSFAC PATHWAYS package includes all components of the TRANSFAC BASIC package, and extends its capabilities with TRANSPATH®—a comprehensive database of mammalian signal transduction and metabolic pathways—along with the Pathfinder tool and the Pathway Omics Suite.
TRANSPATH database
TRANSPATH® is a database of mammalian signal transduction and metabolic pathways. As one of the earliest pathway databases ever created, it has grown since to the remarkable volume of more than 1,244,392 manually curated reactions. One of the largest pathway databases available, optimally suited for geneXplain’s proprietary Upstream Analysis.
Structure
TRANSPATH organizes the information about genes/molecules and reactions according to multiple hierarchies. Its sophisticated structure makes it one of the scientifically best conceptualized pathway resources, suitable for multi-purpose uses. It is complemented by one of the richest corpora of pathway data available among all public domain and commercial sources, all manually curated by experts.
Reaction hierarchy in the TRANSPATH® database on molecular pathways:
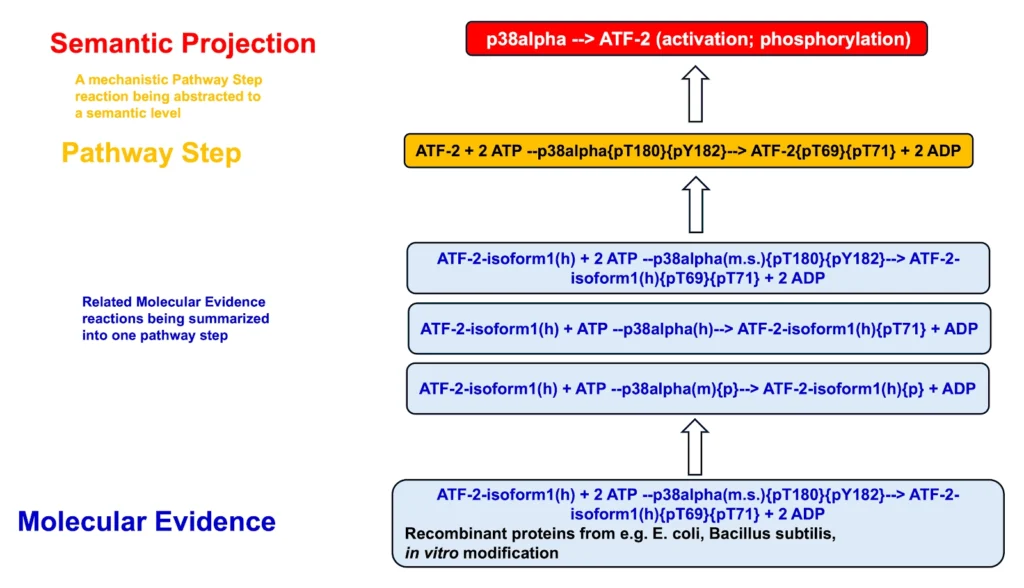
Individual reactions are documented with all experimental details, in a strictly mechanistic way that includes all reaction partners and the taxonomic origin of each molecule as reported in the published experiment (“molecular evidence level”). All evidence for a certain pathway step is accumulated to provide a more comprehensive and complete picture (“pathway step level”). On top, a semantic view is provided, which focuses on the key components only and omits mechanistic details as well as small abundant molecules (“semantic projection”). Complete networks and pathways are built from molecules and their reactions.
To consider the heterogeneity of information given in the original publications, TRANSPATH transparently but precisely differentiates protein molecules according to:
Their relatedness within one genome
Information can be specifically retrieved regarding:
(a) specific individual proteins,
(b) all products of a certain gene (isoforms),
(c) different family relation levels (e.g., paralogs);
Their relatedness between different genomes (orthology)
Their association and modification status(a) protein complexes are specified with their exact composition;
(b) post-translational modifications are given with their exact positions in the protein.
TRANSPATH Pathways
TRANSPATH database collects and systematizes canonical signal transduction and metabolic pathways. Currently it collects over 1500 various pathways.
Pathway visualisation
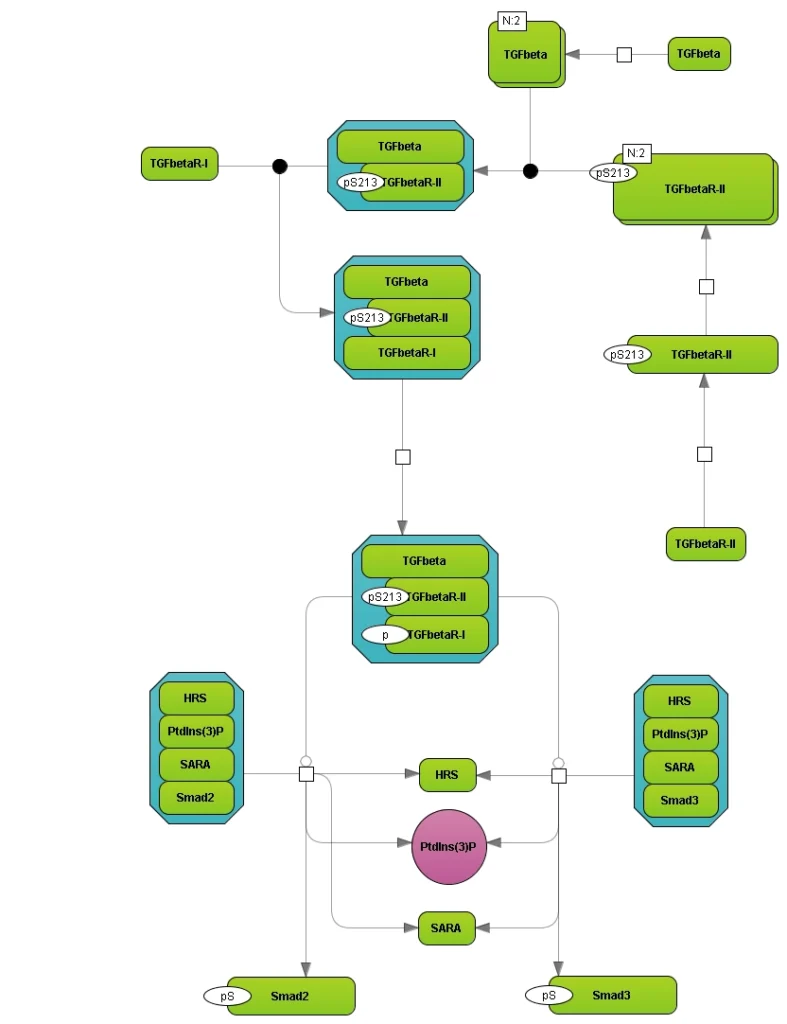
We record precise information about molecular details of the pathway reactions including posttranslational modifications of the molecules involved, their complexes and isoforms.
Visualization of the TGFbeta network with the geneXplain platform; data were retrieved from the TRANSPATH® database.

For each of the pathways contained in TRANSPATH database, users find explicit information on the pathway subcomponents and individual reactions, from which the pathway subcomponents are assembled. Below is a fragment of the detailed description of the IFNalpha/beta pathway.
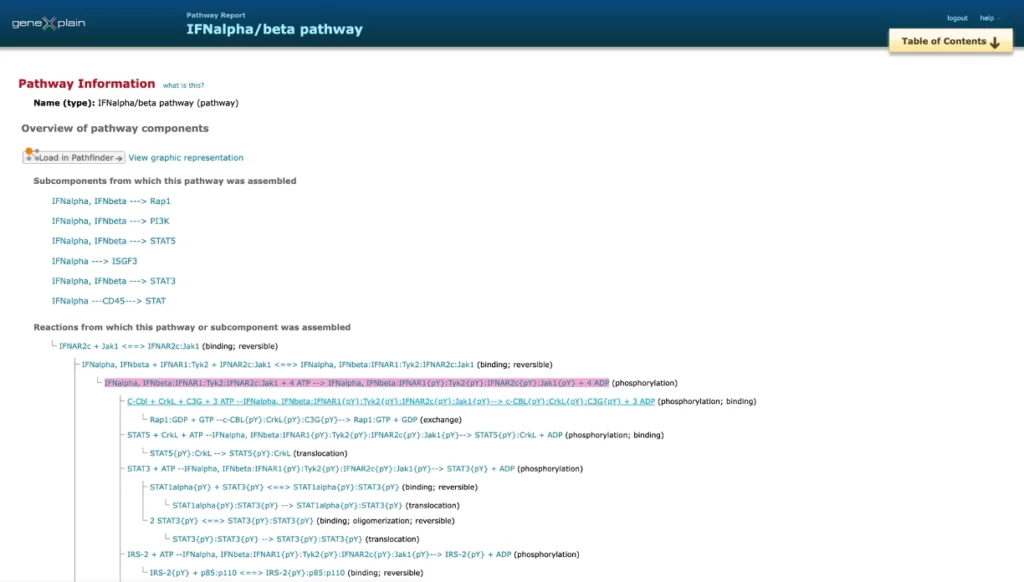
Individual reactions are documented with all experimental details, in a strictly mechanistic way that includes all reaction partners and the taxonomic origin of each molecule as reported in the published experiment (“molecular evidence level”).
All evidence for a certain pathway step is accumulated to provide a more comprehensive and complete picture (“pathway step level”). On top, a semantic view is provided, which focuses on the key components only and omits mechanistic details as well as small abundant molecules (“semantic projection”).
Complete networks and pathways are built from molecules and their reactions. It collects over 1,200,000 experimentally proven reactions of phosphorylation, acetylation, ubiquitination, translocation and other types of reactions involved in signal transduction. These reactions build a highly connected regulatory reference network which is used for analysis and reconstruction of molecular mechanisms of diseases, identification of master-regulators and drug targets.
Pathways analysis
The collected signal transduction networks can be used by the included tools for pathway analysis, particularly, in connection with TF-DNA binding motifs, this allows upstream analysis as integrated promoter-network analysis, whereby the TFs of in step 1 predicted TFBSs are used as starting point for finding master regulators in step 2 converging upstream of the transcription factors. Key nodes where these pathways converge are considered as master regulators of the process under study.
Activities of transcription factors (TFs, blue circles) are regulated by upstream signaling cascades (components shown as green circles). These converge in potential master regulators of the process under study.
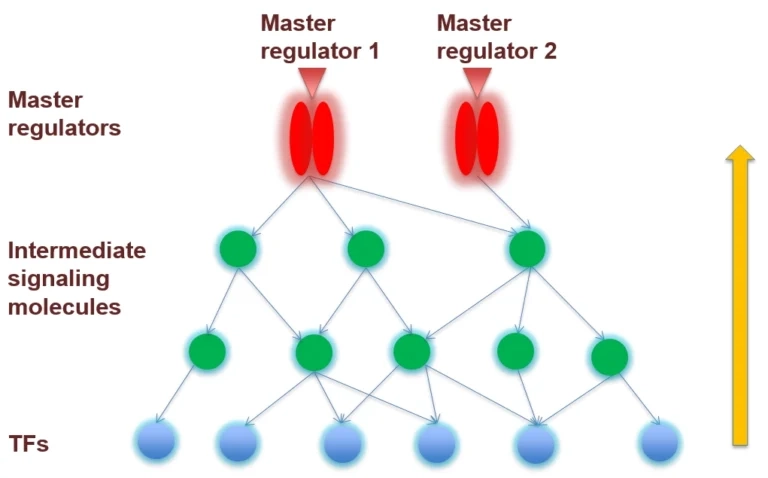
Reaction network
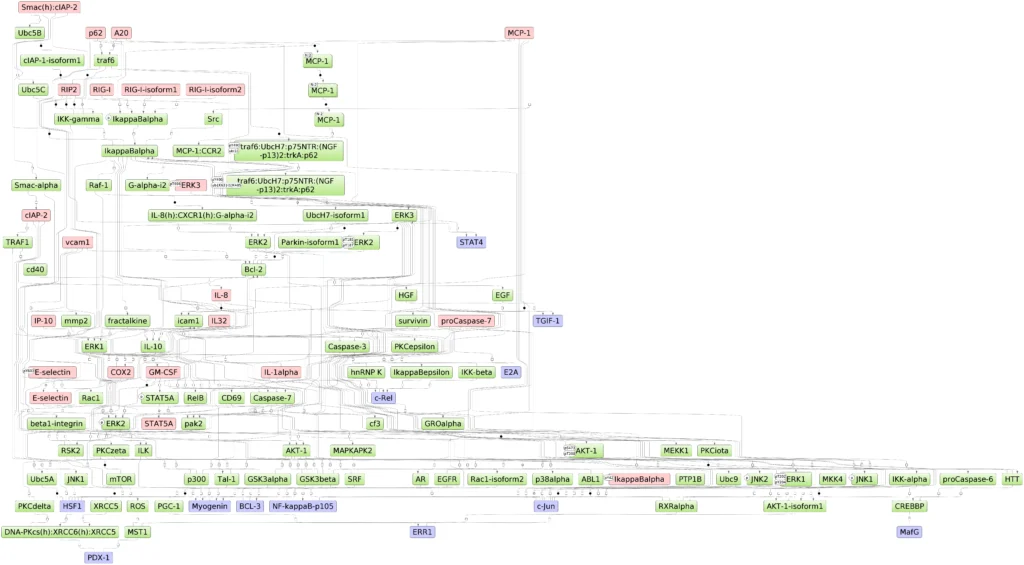
TRANSPATH® database is one of the most comprehensive repositories of signal transduction reactions in mammalian cells. It collects over 1,200,000 experimentally proven reactions of phosphorylation, acetylation, ubiquitination, translocation and other types of reactions involved in signal transduction. These reactions build a highly connected regulatory reference network which is used for analysis and reconstruction of molecular mechanisms of diseases, identification of master-regulators and drug targets.
TRANSPATH® reaction reference network (a fragment)
A fragment of TRANSPATH(R) reaction reference network. Pink nodes correspond to the “master-regulators”, and blue nodes correspod to “effectors”.
Upstream analysis
The TRANSFAC PATHWAYS package uniquely integrates promoter analysis with pathway analysis, enabling the systematic identification of master regulators within gene regulatory networks. This level of combined regulatory and pathway insight is not available in other tools on the market.
What is Upstream Analysis?
GeneXplain’s proprietary approach to analyze gene expression data is called Upstream Analysis. The term indicates that it is a causal analysis, providing a clue about the reason why a certain set of genes has been up- (or down-) regulated in the system under study. In contrast, conventional analyses usually reveal the effects of the differentially expressed genes, e.g. by mapping them onto ontological categories.
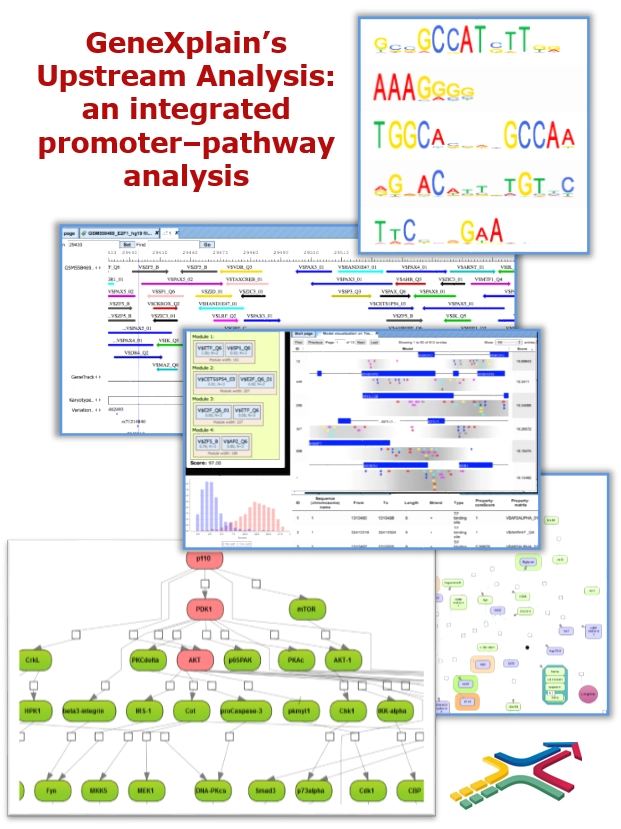
How does it work?
GeneXplain’s Upstream Analysis is an integrated promoter – pathway analysis. It starts from any list of differentially expressed genes (DEGs), which you may have extracted from your raw data with the aid of the geneXplain platform, and comprises two main steps:
- At first, the promoters of the differentially regulated genes are retrieved and analyzed for potential transcription factor (TF) binding sites and their combinations. From that, a set of TFs is identified that potentially have regulated the found DEGs.
- In a second step, the pathways are reconstructed that are known to activate the previously hypothesized TFs. Molecules where these pathways converge are considered as potential master regulators of the process under study
Step 1: Promoter analysis
First, potential transcription factor binding sites (TFBSs) are identified in all promoters of the DEGs of your experiment (Yes set) as well as in a negative control set (No set). This is usually done with a library of position-specific scoring or positional weight matrices (PSSMs or PWMs).
We recommend applying the most comprehensive matrix library available, the TRANSFAC® database, and using the MATCHTM algorithm for the sequence analysis.
Next, out of all these potential transcription factor binding sites (TFBSs), those that are characteristic for the DEG set under study are identified. This is done by rigorously determining their enrichment in the Yes- compared to the No set.
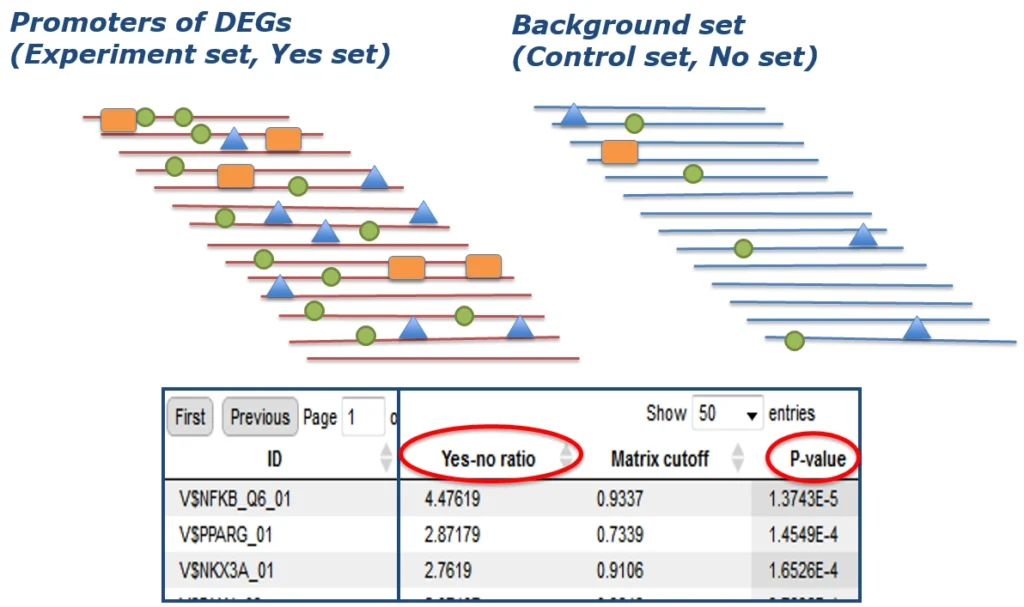
Step 2: Pathway analysis
Step 1 resulted in a set of transcription factors (TFs), that are likely responsible for the differential regulation of the observed set of DEGs. From available pathway data, we have extracted information about all relevant signaling cascades that regulate the activity of TFs; optimally, the TRANSPATH(R) database is used for this and the further analysis.
As has been proven in a large number of use cases, these pathways usually converge in a couple of key nodes, which qualify as candidate master regulators of the process under study.

Detailed Upstream analysis- multi-omics workflow
TRANSFAC® Pathways supports end-to-end multi-omics analysis—from data upload and integration to comprehensive analysis across genomics, transcriptomics, epigenomics, proteomics, and metabolomics.
Modern omics technologies routinely deliver long lists of differentially expressed genes, methylation changes, or protein abundance shifts. However, these downstream readouts do not immediately explain which signaling cascades are responsible for the observed changes, or which upstream regulators should be targeted to modulate the system.
The upstream of TFs concept closes this gap. Starting from transcription factors (TFs) inferred from promoter/enhancer analysis, we trace the regulatory architecture backwards through signaling networks to the receptors, kinases, and other key nodes that drive these TFs. This reveals causal links between:
- disease- or phenotype-specific gene expression signatures,
- the transcription factors controlling those signatures, and
- the signaling pathways and master regulators that act upstream of these TFs.
This “from TFs upstream to signaling” strategy is at the core of geneXplain’s pathway-centric analysis and provides a powerful bridge between regulatory genomics and systems-level signaling biology.
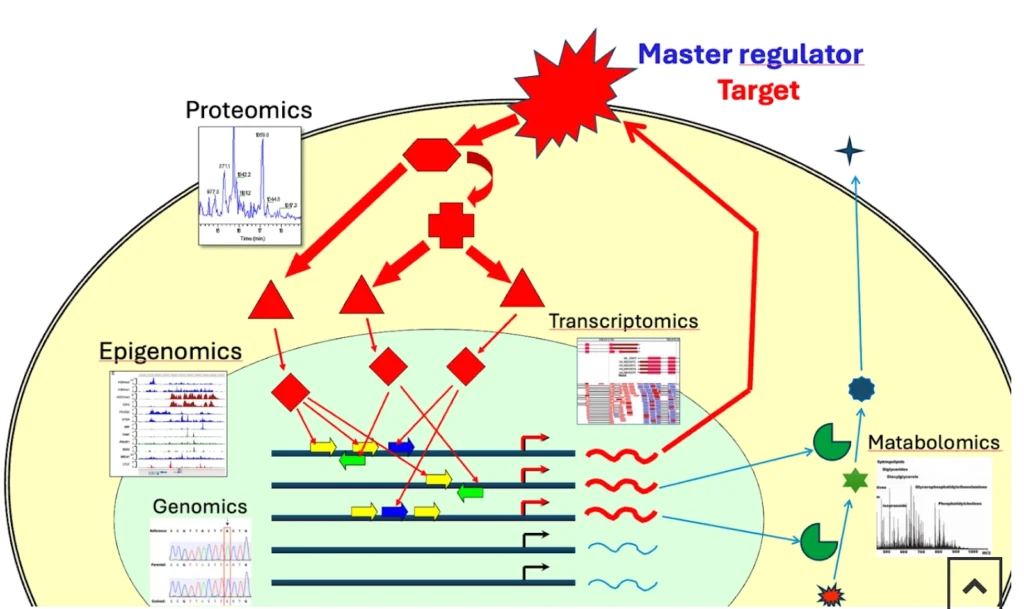
From gene expression changes to signaling cascades
Typical omics workflows start with one or more of the following layers:
- Transcriptomics – differentially expressed genes (DEGs) describing the phenotype or pathology.
- Epigenomics – changes in regulatory regions (e.g. ChIP-seq peaks, DNA methylation) that impact TF binding.
- Genomics – variants in promoters, enhancers, or coding regions that affect regulation.
- Proteomics / phosphoproteomics – altered abundance or phosphorylation of signaling proteins and TFs.
- Metabolomics – end-point readouts of pathway activity that help to focus on the most functionally critical DEGs.
By integrating these layers, we obtain a robust gene set characterizing the condition of interest. The key question then becomes:
Which transcription factors and upstream signaling cascades are responsible for this gene set?
Answering this question requires connecting regulatory regions (where TFs act) with signaling pathways (where extracellular and intracellular signals are processed).
Key steps in upstream-of-TF analysis
Step 1 – Constructing the gene set describing the process
The analysis begins by constructing a gene set that best represents the studied biological process or pathology:
- From transcriptomics, differential expression or clustering identifies up- and down-regulated genes.
- From epigenomics, peak-to-gene or CpG-to-gene mapping connects regulatory changes to nearby genes.
- From proteomics, protein identifiers are converted to corresponding genes.
- From genomics, variants are mapped to affected genes, especially when they fall into regulatory regions.
The result is a set (or several sets) of genes that show coordinated behavior in the condition under study and serve as the entry point for regulatory and pathway analysis.
Step 2 – Promoter/enhancer analysis and TF identification
For each gene set, promoter and/or enhancer regions are analyzed to detect overrepresented transcription factor binding sites (TFBSs):
- Regulatory regions are defined (e.g. ±1 kb around transcription start sites, or known enhancer coordinates).
- These regions are scanned with high-quality TF binding models to detect enriched motifs.
- Complexes and modules of co-acting TFs (combinatorial regulation) are identified to reflect realistic regulatory logic.
This step transforms a “flat” gene list into a regulatory profile: a small set of transcription factors and TF modules that are statistically most likely to control the observed gene expression changes.
Step 3 – Mapping TFs onto signaling networks
Once candidate TFs are identified, the analysis proceeds upstream into signaling space:
- TFs are mapped onto a curated signaling and metabolic network.
- Known regulatory relationships (e.g. which kinases activate a TF, which receptors feed into a pathway that converges on that TF) are retrieved.
- Upstream interaction chains are reconstructed, linking TFs to receptors, kinases, phosphatases, adaptor proteins, and other signaling molecules.
This step builds signal transduction routes that explain how external or internal cues propagate through the network and converge on the TFs controlling the gene set.
Step 4 – Upstream network reconstruction and master regulators
The next goal is to identify master regulators – those key upstream nodes whose activity can influence large parts of the network:
- Network algorithms are applied to trace shortest or most plausible signaling paths from TFs back to upstream nodes.
- Nodes are ranked by criteria such as connectivity, frequency of occurrence across reconstructed paths, and their position in the hierarchy (e.g. receptors vs. downstream kinases).
- The result is a compact set of candidate master regulators (receptors, kinases, TFs, other signaling proteins) that function as control points for the observed transcriptional program.
These master regulators provide a mechanistic explanation of the phenotype and serve as high-value hypotheses for further experimental validation.
Step 5 – Pathway and process interpretation
With TFs and upstream regulators defined, the analysis moves to biological interpretation:
- The identified regulators and their downstream targets are mapped to signaling and metabolic pathways.
- Enrichment analysis highlights pathways and processes most consistently affected in the condition.
- Integrated visualization shows how signals flow from receptors, through signaling cascades, to TFs and finally to target genes.
This provides a systems-level view that connects:
- the original omics readouts (e.g. DEGs),
- the TFs inferred from regulatory region analysis, and
- the upstream signaling modules that explain those TF activities.
What you gain from upstream-of-TF analysis
Applying the upstream-of-TF approach allows you to:
- Move from long lists of altered genes to a small set of mechanistically justified regulators.
- Link transcription factor activity directly to concrete signaling cascades and pathways.
- Identify master regulators that act as bottlenecks or control points in the network.
- Generate focused, experimentally testable hypotheses on how to modulate the system (e.g. by targeting specific receptors, kinases, or TFs).
- Use multi-omics evidence to prioritize the most robust and biologically relevant regulators.
This upstream-of-TF strategy is implemented in geneXplain’s analysis environment, where promoter/enhancer analysis, TF motif enrichment, and signaling-network reconstruction are integrated into a coherent workflow, providing a clear mechanistic bridge from transcription factors to signaling cascades.
PathFinder
PathFinder is the tool for visualization of signaling and metabolic pathways using (System Biology Graphic Notation) SBGN standard. It is an integrative part of the TRANSPATH database.
The PathFinder tab automatically opens in your browser after you have selected to visualize any of the pathways in the geneXplain portal interface by clicking on the PathFinder link:
Upon opening any pathway from the geneXplain portal interface in the PathFinder, you will see two diagrams:
One will be automatically layouted in respect to the intracellular compartments its elements belong to:
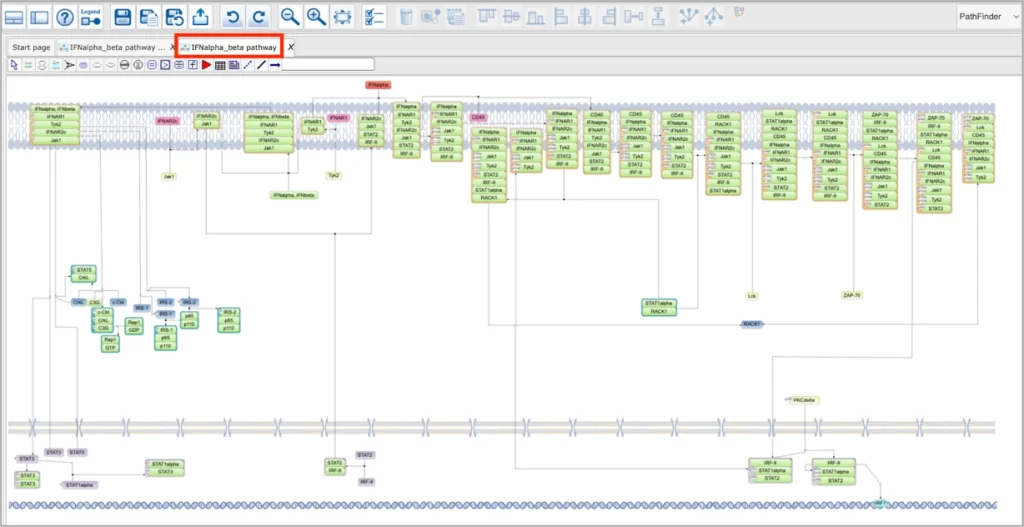
And the other one will not contain compartments and will be subject to user-selected layout in the tools menu:
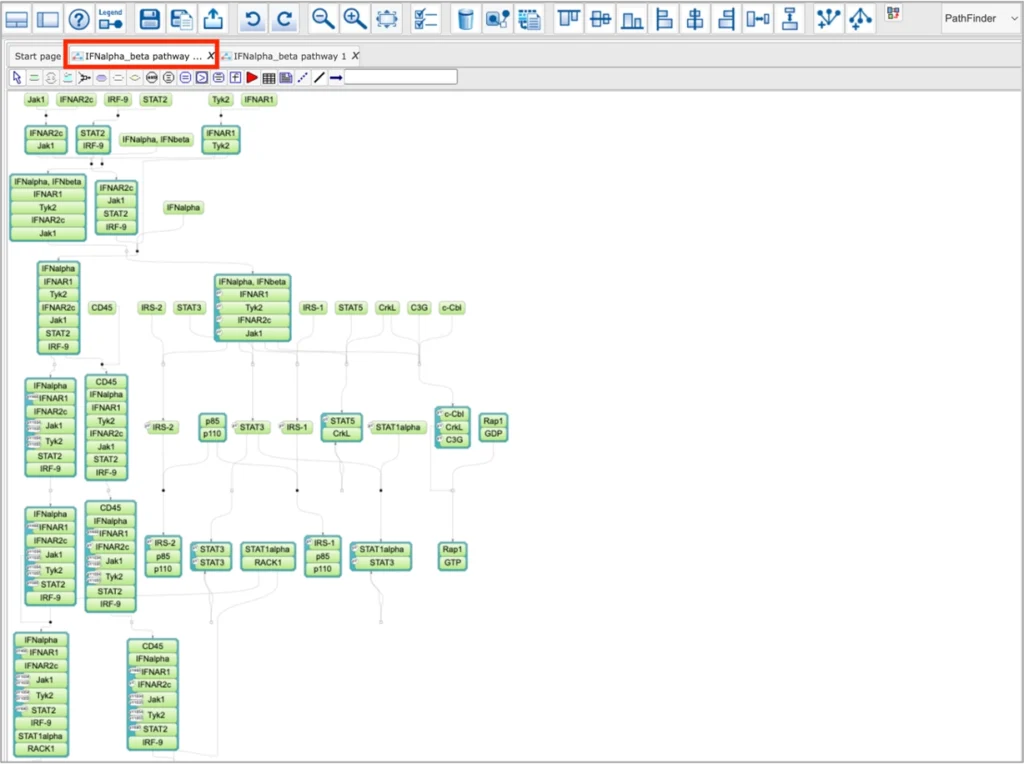
All PathFinder diagrams without compartments have a standard SBGN and SBML operations menu (toolbar) available on the top of the diagram:

You will find further details on the PathFinder tool in its User Guide.
ODE Modeling in TRANSFAC PATHWAYS
The TRANSFAC PATHWAYS package integrates advanced modeling tools within the geneXplain platform, built on the BioUML simulation environment. According to an independent benchmark study (Maggioli et al., Bioinformatics, 2019), BioUML is recognized as one of the most efficient SBML simulation engines available, demonstrating superior speed compared to alternatives such as COPASI, SystemModeler, and SBML2Modelica across 613 curated models from the BioModels database.
Notably, BioUML is the only platform that successfully passes 100% of simulation tests in the SBML Test Suite Core v3.3.0. Its simulation engine supports a wide range of mathematical formalisms, including:
- Ordinary Differential Equations (ODE)
- Differential-Algebraic Equations (DAE)
- Hybrid models
- One-dimensional Partial Differential Equations (1D PDE)
Beyond simulation, BioUML provides a comprehensive environment for visual and systems-level modeling, enabling:
- Network modeling
- Composite (multi-scale) modeling
- Agent-based modeling
The intuitive web-based graphical user interface (GUI) allows researchers to construct and explore complex biological models efficiently.
Parameter Fitting and Optimization
BioUML includes a powerful parameter fitting engine designed to support diverse experimental scenarios:
- Fitting to time-course or steady-state data
- Multi-experiment parameter fitting
- Constraint-based optimization
- Handling of local and global parameters
- Custom optimization workflows via JavaScript
A variety of advanced optimization algorithms are implemented, including:
- Adaptive simulated annealing
- Evolutionary programming
- Particle swarm optimization
- Stochastic ranking evolution strategy
- Cellular genetic algorithms
These capabilities enable precise calibration of biological models and facilitate robust simulation of complex regulatory and signaling processes.
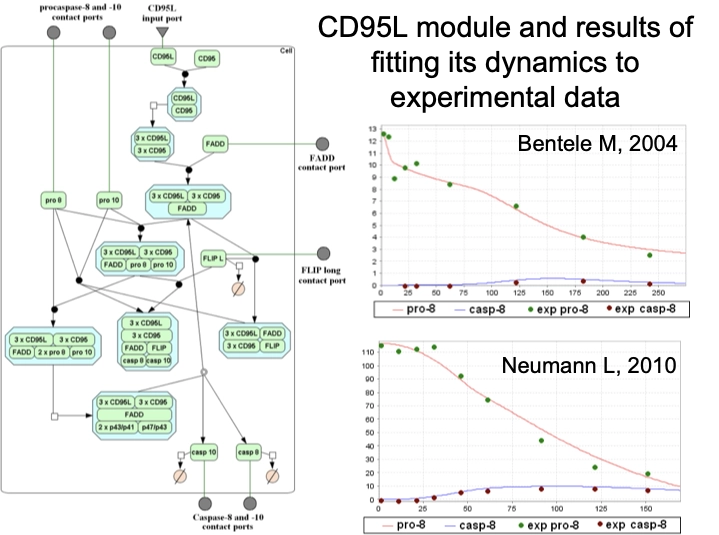
Application Example
An example of these capabilities is demonstrated through an integrated apoptosis model constructed using TRANSFAC PATHWAYS, showcasing how mechanistic pathway knowledge and quantitative modeling can be combined to study disease-relevant processes.
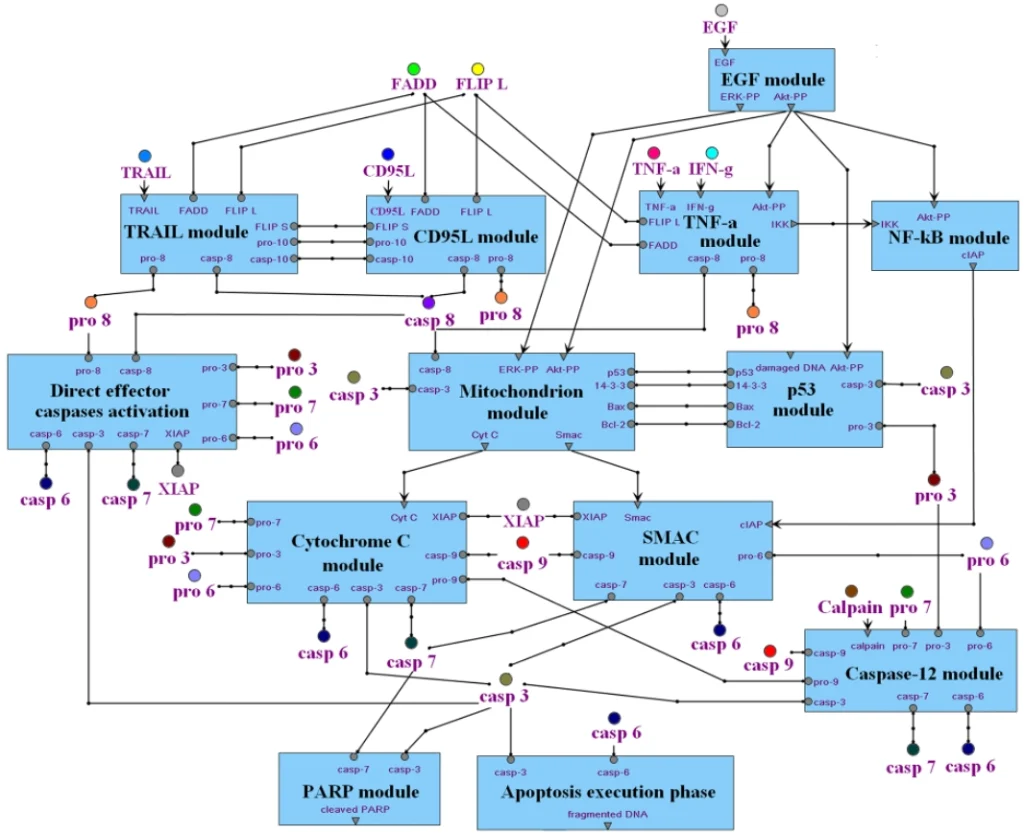
Videos
Browsing pathways in TRANSPATH®+HumanPSD
Functional analysis tool in TRANSPATH®+HumanPSD
Locus report in TRANSPATH®+HumanPSD
TRANSPATH Applications in Research — Selected Publications
Novikova S., Tolstova T., Kurbatov L., Farafonova T., Tikhonova O., Soloveva N., Rusanov A., Zgoda V. (2024) Systems Biology for Drug Target Discovery in Acute Myeloid Leukemia. Int. J. Mol. Sci. 25(9), 4618 (2024) Link
Kisakol, B., Matveeva, A., Salvucci, M., Kel, A., McDonough, E., Ginty, F., Longley, D. B., & Prehn, J. H. M. (2024). Identification of unique rectal cancer-specific subtypes. British journal of cancer, 130(11), 1809–1818. Link
Ivanov, S. M., Tarasova, O. A., & Poroikov, V. V. (2023). Transcriptome-based analysis of human peripheral blood reveals regulators of immune response in different viral infections. Frontiers in immunology, 14, 1199482. Link
Bertram, H., Wilhelmi, S., Rajavel, A., Boelhauve, M., Wittmann, M., Ramzan, F., Schmitt, A. O., & Gültas, M. (2023). Comparative Investigation of Coincident Single Nucleotide Polymorphisms Underlying Avian Influenza Viruses in Chickens and Ducks. Biology, 12(7), 969. Link
Rajavel A., Klees S., Hui Y., Schmitt A.O., Gültas M. (2022) Deciphering the Molecular Mechanism Underlying African Animal Trypanosomiasis by Means of the 1000 Bull Genomes Project Genomic Dataset. Biology (Basel). 11(5), 742. Link
Menck K., Wlochowitz D., Wachter A., Conradi L.C., Wolff A., Scheel A.H., Korf U., Wiemann S., Schildhaus H.U., Bohnenberger H., Wingender E., Pukrop T., Homayounfar K., Beißbarth T., Bleckmann A. (2022) High-Throughput Profiling of Colorectal Cancer Liver Metastases Reveals Intra- and Inter-Patient Heterogeneity in the EGFR and WNT Pathways Associated with Clinical Outcome. Cancers 14(9), 2084. Link
Kechin A.A., Ivanov A.A., Kel A.E., Kalmykov A.S., Oskorbin I.P., Boyarskikh U.A., Kharpov E.A., Bakharev S.Y., Oskina N.A., Samuilenkova O.V., Vikhlyanov I.V., Kushlinskii N.E., Filipenko M.L. (2022) Prediction of EVT6-NTRK3-Dependent Papillary Thyroid Cancer Using Minor Expression Profile. Bull Exp Biol Med. 173(2),252-256. Link
Myer, P. A., Kim, H., Blümel, A. M., Finnegan, E., Kel, A., Thompson, T. V., Greally, J. M., Prehn, J. H., O’Connor, D. P., Friedman, R. A., Floratos, A., & Das, S. (2022). Master Transcription Regulators and Transcription Factors Regulate Immune-Associated Differences Between Patients of African and European Ancestry With Colorectal Cancer. Gastro Hep Adv. 1(3),328-341. Link
Chereda, H., Bleckmann, A., Menck, K., Perera-Bel, J., Stegmaier, P., Auer, F., Kramer, F., Leha, A., & Beißbarth, T. (2021). Explaining decisions of graph convolutional neural networks: patient-specific molecular subnetworks responsible for metastasis prediction in breast cancer. Genome Med. 13(1),42. Link
Menck, K., Heinrichs, S., Wlochowitz, D., Sitte, M., Noeding, H., Janshoff, A., Treiber, H., Ruhwedel, T., Schatlo, B., von der Brelie, C., Wiemann, S., Pukrop, T., Beißbarth, T., Binder, C., & Bleckmann, A. (2021). WNT11/ROR2 signaling is associated with tumor invasion and poor survival in breast cancer. Journal of experimental & clinical cancer research : CR, 40(1), 395. Link
Kalya M., Kel A., Wlochowitz D., Wingender E., Beißbarth T. (2021) IGFBP2 Is a Potential Master Regulator Driving the Dysregulated Gene Network Responsible for Short Survival in Glioblastoma Multiforme. Front Genet. 12, 670240. Link
Benjamin, S.J., Hawley, K.L., Vera-Licona, P., La Vake, C.J., Cervantes, J.L., Ruan, Y., Radolf, J.D., Salazar, J.C. (2021) Macrophage mediated recognition and clearance of Borrelia burgdorferi elicits MyD88-dependent and -independent phagosomal signals that contribute to phagocytosis and inflammation. BMC Immunol. 22, 32. Link
Ivanov, S., Filimonov, D., & Tarasova, O. (2021) A computational analysis of transcriptional profiles from CD8(+) T lymphocytes reveals potential mechanisms of HIV/AIDS control and progression. Comput Struct Biotechnol J. 19, 2447–2459. Link
Meier, T., Timm, M., Montani, M., Wilkens, L. (2021) Gene networks and transcriptional regulators associated with liver cancer development and progression. BMC Med. Genomics 14, 41. Link
Andreev-Andrievskiy, A. A., Zinovkin, R. A., Mashkin, M. A., Frolova, O. Y., Kazaishvili, Y. G., Scherbakova, V. S., Rudoy, B. A., & Nesterenko, V. G. (2021). Gene Expression Pattern of Peyer’s Patch Lymphocytes Exposed to Kagocel Suggests Pattern-Recognition Receptors Mediate Its Action. Frontiers in pharmacology, 12, 679511. Link
Lloyd K., Papoutsopoulou S., Smith E., Stegmaier P., Bergey F., Morris L., Kittner M., England H., Spiller D., White M.H.R., Duckworth C.A., Campbell B.J., Poroikov V., Martins Dos Santos V.A.P., Kel A., Muller W., Pritchard D.M., Probert C., Burkitt M.D.; SysmedIBD Consortium. Using systems medicine to identify a therapeutic agent with potential for repurposing in inflammatory bowel disease. Dis Model Mech. 13(11), dmm044040. Link
Ramzan, F., Klees, S., Schmitt, A. O., Cavero, D., & Gültas, M. (2020) Identification of Age-Specific and Common Key Regulatory Mechanisms Governing Eggshell Strength in Chicken Using Random Forests. Genes (Basel). 11(4), 464. Link
Ayyildiz D., Antoniali G., D’Ambrosio C., Mangiapane G., Dalla E., Scaloni A., Tell G., Piazza S. (2020) Architecture of The Human Ape1 Interactome Defines Novel Cancers Signatures. Sci Rep. 10, 28. Link
Mekonnen, Y.A., Gültas, M., Effa, K., Hanotte, O., Schmitt, A.O. (2019) Identification of Candidate Signature Genes and Key Regulators Associated With Trypanotolerance in the Sheko Breed. Front. Genet. 10, 1095. Link
Nobis, C. C., Dubeau Laramée, G., Kervezee, L., Maurice De Sousa, D., Labrecque, N., & Cermakian, N. (2019) The circadian clock of CD8 T cells modulates their early response to vaccination and the rhythmicity of related signaling pathways. Proc Natl Acad Sci U S A. 116(40), 20077–20086. Link
Orekhov A.N., Oishi Y., Nikiforov N.G., Zhelankin A.V., Dubrovsky L., Sobenin I.A., Kel A., Stelmashenko D., Makeev V.J., Foxx K., Jin X., Kruth H.S., Bukrinsky M. (2018) Modified LDL Particles Activate Inflammatory Pathways in Monocyte-derived Macrophages: Transcriptome Analysis. Curr Pharm Des. 24(26),3143-3151. Link
Wlochowitz, D., Haubrock, M., Arackal, J., Bleckmann, A., Wolff, A., Beißbarth, T., Wingender, E., Gültas, M. (2016) Computational Identification of Key Regulators in Two Different Colorectal Cancer Cell Lines. Front. Genet. 7, 42. Link
Kural, K.C., Tandon, N., Skoblov, M., Kel-Margoulis, O.V. and Baranova, A.V. (2016) Pathways of aging: comparative analysis of gene signatures in replicative senescence and stress induced premature senescence. BMC Genomics 17(Suppl 14), 1030. Link
Malusa, F., Taranta, M., Zaki, N., Cinti, C., & Capobianco, E. (2015) Time-course gene profiling and networks in demethylated retinoblastoma cell line. Oncotarget. 6(27), 23688–23707. Link
Kutumova E.O., Kiselev I.N., Sharipov R.N., Lavrik I.N., Kolpakov F.A. (2012) A modular model of the apoptosis machinery. Adv Exp Med Biol. 736, 235-45. Link
Schuler, M., Keller, A., Backes, C., Philippar, K., Lenhof, H. P., & Bauer, P. (2011) Transcriptome analysis by GeneTrail revealed regulation of functional categories in response to alterations of iron homeostasis in Arabidopsis thaliana. BMC Plant Biol. 11, 87. Link
Ante M., Wingender E., Fuchs M. (2011) Integration of gene expression data with prior knowledge for network analysis and validation. BMC Res Notes. 4,520. Link
Chiu SC, Tsao SW, Hwang PI, Vanisree S, Chen YA, Yang NS. (2010) Differential functional genomic effects of anti-inflammatory phytocompounds on immune signaling. BMC Genomics. 11, 513. Link
How to cite TRANSPATH
Krull M, Pistor S, Voss N, Kel A, Reuter I, Kronenberg D, Michael H, Schwarzer K, Potapov A, Choi C, Kel-Margoulis O, Wingender E. TRANSPATH: an information resource for storing and visualizing signaling pathways and their pathological aberrations. Nucleic Acids Res. 2006;34(90001):D546-D551. Link