TRANSFAC 2.0 — The Most Comprehensive Transcription Factor Resource
Explore TRANSFAC packages
About TRANSFAC BASIC
What is included:
Database Features
Tool Features
Statistics
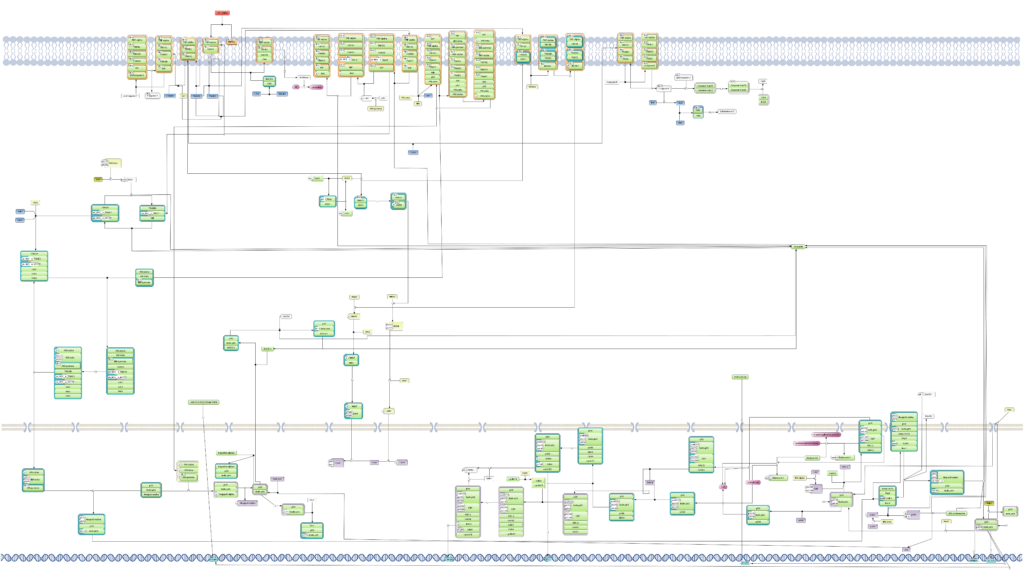
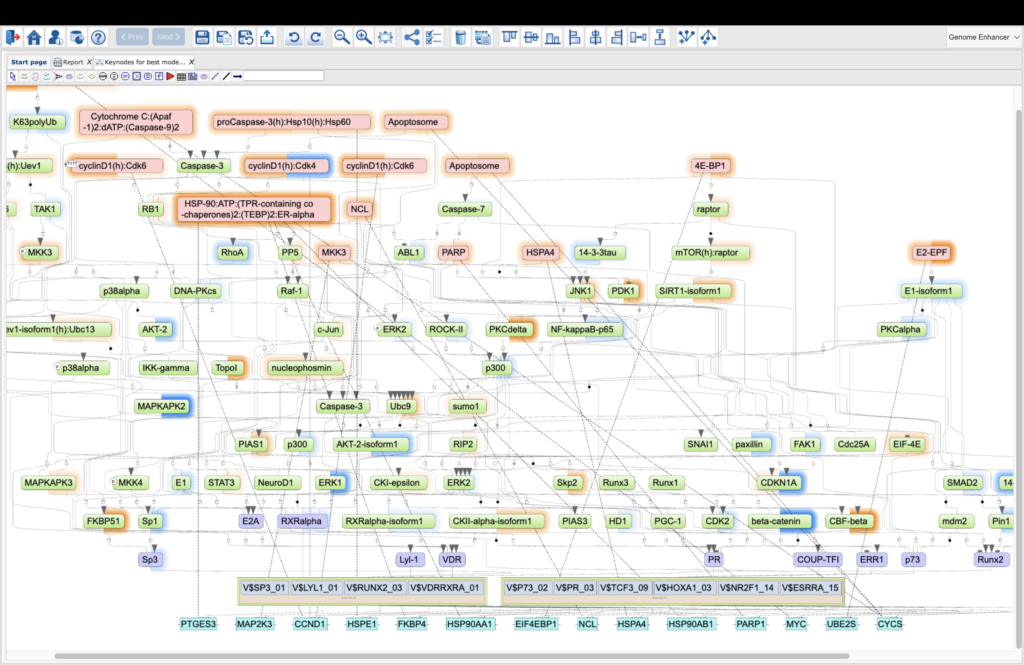
About TRANSFAC PATHWAYS
What is included:
Database Features
Tool Features
Statistics
About TRANSFAC DISEASES
What is included:
Database Features
Tool Features
Statistics
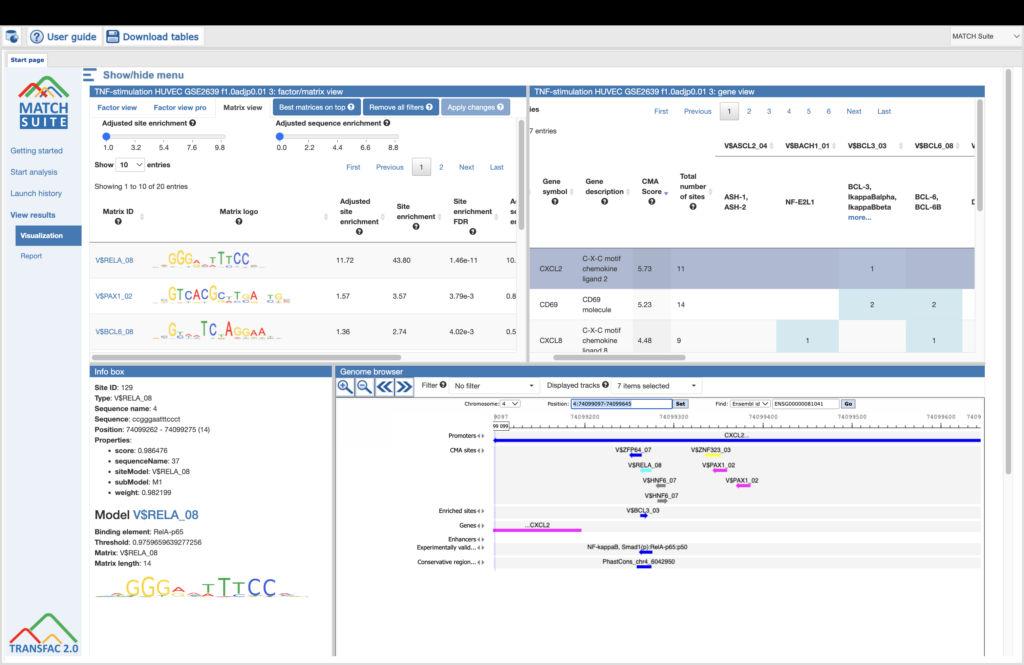
TRANSFAC is the gold-standard database of eukaryotic transcription factors (TFs), their binding sites, and DNA binding models — manually curated, experimentally validated, and integrated with powerful analysis tools.
Whether you’re exploring gene regulation, reconstructing signaling networks, or hunting disease biomarkers, the TRANSFAC suite gives you the knowledge base and analysis tools you need.