Deyneko, I. V., Kel, A. E., Kel-Margoulis, O. V., Deineko, E. V., Wingender, E. and Weiss, S. (2013) MatrixCatch – a novel tool for the recognition of composite regulatory elements in promoters. BMC Bioinformatics 14, 241. Link
Stegmaier, P., Kel, A., Wingender, E., Borlak, J. (2013) A discriminative approach for unsupervised clustering of DNA sequence motifs. PLoS Comput. Biol. 9, e1002958. Link
Stegmaier P., Voss N., Meier T., Kel A., Wingender E., Borlak J. (2011) Advanced computational biology methods identify molecular switches for malignancy in an EGF mouse model of liver cancer. PLoS ONE 6, e17738. PubMed.
Wingender E. (2008) The TRANSFAC project as an example of framework technology that supports the analysis of genomic regulation. Brief Bioinform. 9:326-332. PubMed.
Kel A., Konovalova T., Waleev T., Cheremushkin E., Kel-Margoulis O., Wingender, E. (2006) Composite Module Analyst: a fitness-based tool for identification of transcription factor binding site combinations. Bioinformatics 22:1190-1197. PubMed.
Waleev T. Shtokalo D., Konovalova T., Voss N., Cheremushkin E., Stegmaier P., Kel-Margoulis O., Wingender E., Kel A. (2006) Composite Module Analyst: identification of transcription factor binding site combinations using genetic algorithm. Nucleic Acids Res. 34, W541-W545. PubMed.
Kel A.E., Gössling E., Reuter I., Cheremushkin E., Kel-Margoulis O.V., Wingender E. (2003) MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 31:3576-3579. PubMed
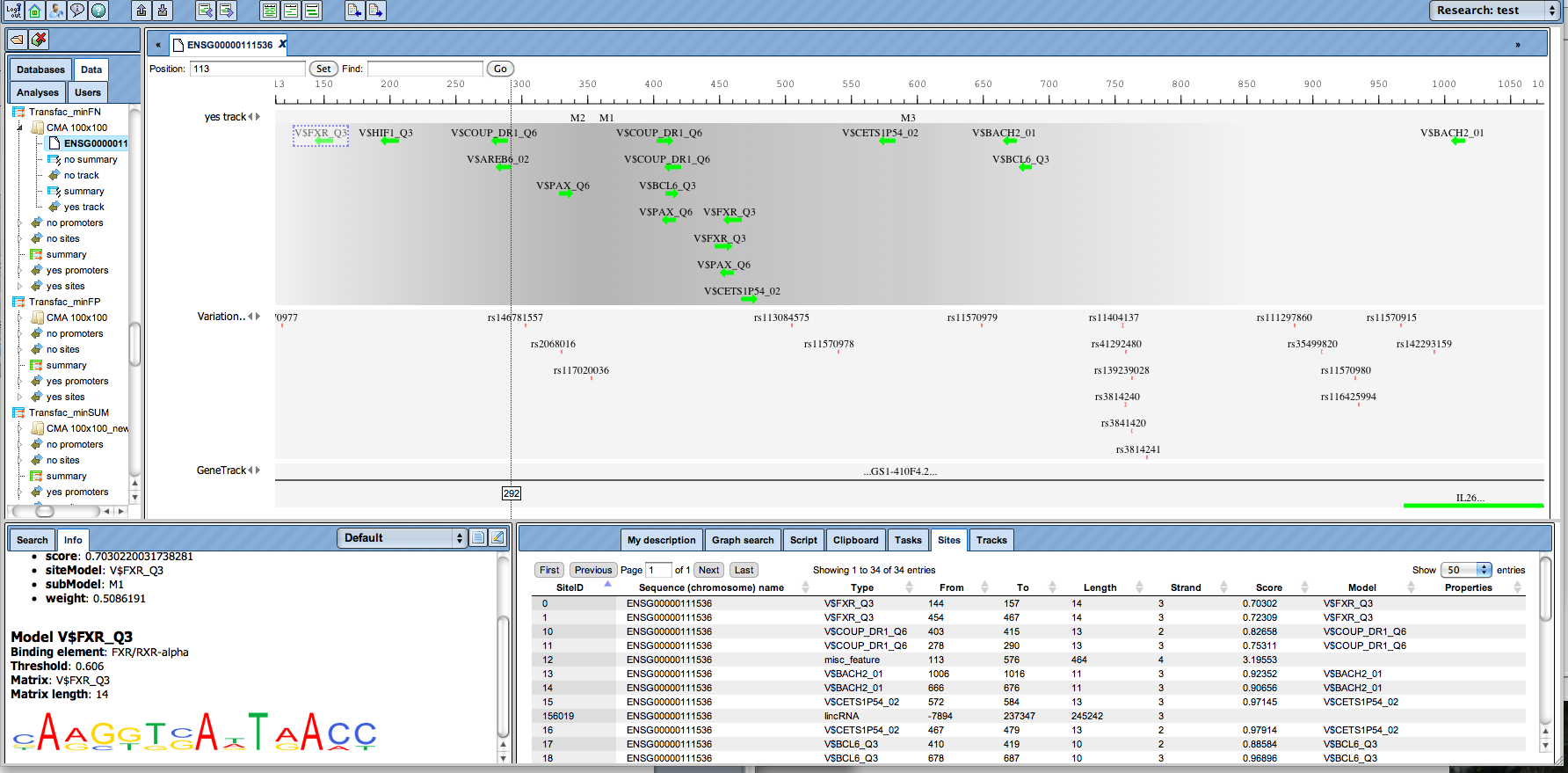
Get the most comprehensive overview about potential regulatory elements in your sequences.
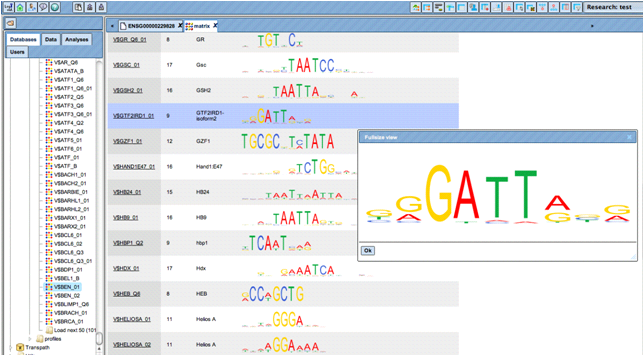
Fast and immediate analysis with a small library of positional weight matrices.
Make use of the full potential of the gold standard in the field, the TRANSFAC® database.
Automatically generate complex promoter models.
Generate high-quality hypotheses about the transcription factors acting on a given set of co-regulated promoters.
Search for additional genes fitting to your promoter model.