Release 2025.2 Announcement
The geneXplain team is proud to present the new release of its products:
- TRANSFAC®,
- TRANSPATH® and HumanPSD™ databases release 2025.2,
- geneXplain® platform release 7.7 and Genome Enhancer release 3.7,
All coming in respective TRANSFAC 2.0 packages.
With this release we are especially proud to present significant developments in the TRANSFAC Pathways package.
TRANSFAC PATHWAYS
(Featuring TRANSFAC® 2.0 2025.2, TRANSPATH® 2025.2, the geneXplain® platform 7.7, and the Pathway Omics Suite 3.6)
Pathway Omics Suite 3.6— New End-to-End Multi-Omics Workflow
One of the highlights of the 2025.2 release is an introduction of the Pathway Omics Suite 1.0, an integrated pipeline designed to transform raw multi-omics datasets into clear, causal biological mechanisms. Designed for biologists, translational researchers, and drug-discovery teams, it reveals master regulators—the key molecules that drive disease-specific regulatory cascades.
Built for both translational research and systems biology, the Suite offers:
- Unified multi-omics integration (Transcriptomics, epigenomics, proteomics, and more)
- Automated differential analysis and molecular feature annotation
- Upstream regulator discovery using curated TRANSFAC® and TRANSPATH® knowledge
- Mechanistic reconstruction of signaling and metabolic pathways
- Publication-ready visualizations (Pathway diagrams, regulator trees, network maps)
- Comprehensive, exportable result reports with full methodological traceability.
With its no-coding interface and the power of curated molecular knowledge, the Pathway Omics Suite accelerates target discovery, biomarker identification, and hypothesis generation—from minutes to insights.
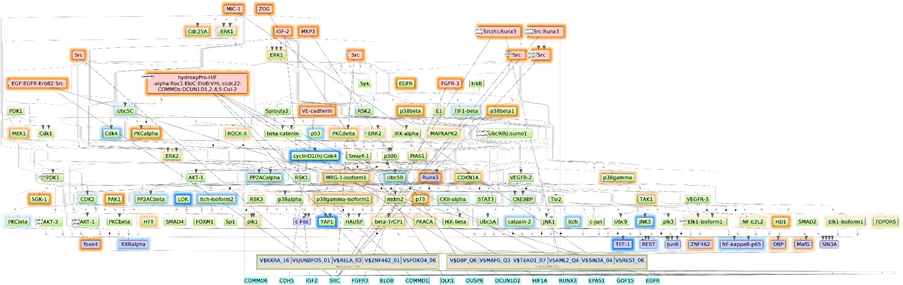
With this release, the Pathway Omics Suite becomes an indispensable part of TRANSFAC Pathways package.
New Database: Protein Genome Map (PGM) 2025.2
This release also marks the launch of a new database: the Protein Genome Map.
The Protein Genome Map is a database of genomic coordinates of protein functional features derived from high quality, manual curation in TRANSPATH as well as additional sources. The focus is on known post-translational modifications. With this novel resource, curated information now becomes available for functional genomics investigations such as variant effect analysis.
The database can be queried using the accompanying method “Create protein feature track”. The method extracts genomic intervals encoding protein functional features that overlap genome coordinates of interest into the geneXplain platform track which can then be exported or further analyzed with a portfolio of platform tools and workflows.
TRANSFAC BASIC
(Featuring TRANSFAC® 2.0 2025.2 and geneXplain® platform 7.7)
The geneXplain® platform tool in its new release 7.7 contains the following new features:
- MuSiC deconvolution package
MuSiC utilizes cell-type specific gene expression from single-cell RNA sequencing (RNA-seq) data to characterize cell type compositions from bulk RNA-seq data in complex tissues. By appropriate weighting of genes showing cross-subject and cross-cell consistency, MuSiC enables the transfer of cell type-specific gene expression information from one dataset to another.
Solid tissues often contain closely related cell types which leads to collinearity. To deal with collinearity, MuSiC employs a tree-guided procedure that recursively zooms in on closely related cell types. Briefly, we first group similar cell types into the same cluster and estimate cluster proportions, then recursively repeat this procedure within each cluster.
- BigWig to BED converter
This tool converts a BigWig file to a BED format and filters values below the threshold.
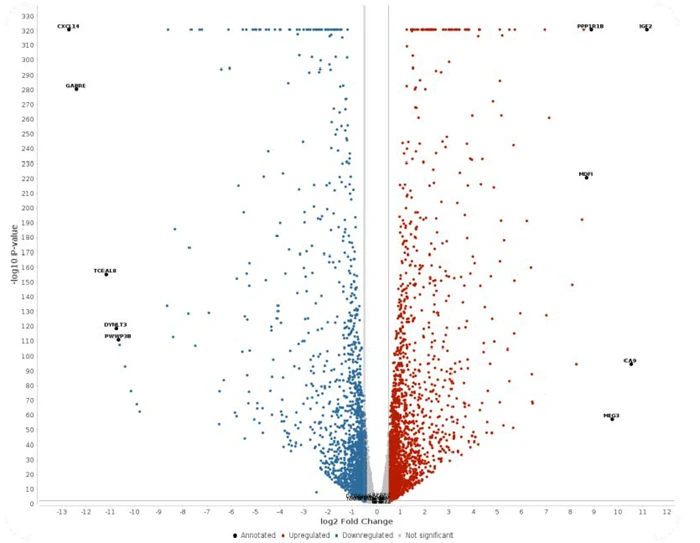
- Volcano plot
A volcano plot is a graphical representation commonly used in differential gene expression studies to simultaneously display the magnitude of expression change and its statistical significance. The x-axis reflects log₂ fold change, indicating the direction and extent of regulation, while the y-axis shows −log₁₀ of the p-value, representing statistical confidence.
This visualization enables rapid identification of differentially expressed genes.
TRANSFAC DISEASES
(Featuring TRANSFAC® 2.0 2025.2, TRANSPATH® 2025.2, HumanPSD™ 2025.2, the geneXplain® platform 7.7 and Genome Enhancer 3.7)
The Genome Enhancer tool in its new release 3.7 contains the following new features:
- Genome Enhancer now includes several new parameters in the Expert settings. Users can customize previously fixed defaults, allowing them to define preferred statistical cut-offs, filtering criteria, the size and number of promoters used for analysis, as well as parameters governing epigenomic and transcriptomic interactions.
- Genome Enhancer now features an enhanced CpG DNA methylation analysis, enabling the integrated assessment of CpG hyper- and hypomethylated regions alongside differentially expressed genes (DEGs).
- All Genome Enhancer demo reports have been updated to Release 3.7, ensuring full compatibility with the latest pipelines, databases, and interface features.
