geneXplain platform
The geneXplain platform provides an online toolbox and workflow management system for a wide range of bioinformatics and systems biology applications, with 100 GB of integrated cloud storage included in every TRANSFAC® package.
Structure
GeneXplain platform has individual modules, or Bricks that are unified under a standardized interface, with a consistent look-and-feel and can flexibly be put together to comprehensive workflows. The workflow management is intuitively handled through a simple drag-and-drop system. With this system, you can edit the predefined workflows or compose your own workflows from scratch.
Own Bricks can easily be added as scripts or plug-ins and can be used in combination with pre-existing analyses.
GeneXplain platform provides a number of state-of-the-art bricks; some of them can be obtained free of charge, while others require licensing for small fee in order to guarantee active maintenance and dynamic adaptation to the rapidly developing know-how in this field.
Key Features
Comprehensive Omics Analysis
Analyze virtually any type of omics data within a single platform, including RNA-seq, ChIP-seq, ATAC-seq, CUT&RUN, WGS, WES, and more—including single-cell datasets.
200+ Integrated Tools and Pipelines
Access over 200 tools for analyzing gene lists, differentially expressed genes (DEGs), and diverse data types across hundreds of input formats (FASTA, FASTQ, VCF, BED, etc.). The platform integrates with Galaxy and provides a flexible workflow management system with ready-to-use pipelines across all areas of bioinformatics.
Intuitive Web-Based Interface (No Coding Required)
Work efficiently through a powerful graphical user interface (GUI) accessible directly in your web browser—no installation or programming skills required. The platform enables fully no-code bioinformatics analysis.
Flexible API for Advanced Workflows
Extend and automate analyses a comprehensive API available for Python, R, and Java. Custom workflows and integrations can be easily implemented.
Access: https://github.com/genexplain
Integrated Cloud Storage
Each TRANSFAC® package includes 100 GB of cloud storage, allowing users to upload data, manage results, and organize analyses across unlimited projects within the geneXplain platform.
Advanced Analytical Capabilities
Integrated AI and ML tools for TFBS prediction
The platform provides access to advanced tools for prediction of genomic transcription factor (TF) binding sites and composite regulatory regions using such algorithms of Machine Learning (ML) and Artificial Intelligence (AI) as Genetic Algorithms and Sparse Logistic Regression.
Integrated databases and analysis tools
The platform provides an integrated view on several databases and analysis tools, public domain as well as commercial ones. They can be combined in a highly flexible way to design customized analyses.
Ready-made workflows for an easy start
A rapidly growing number of proven workflows facilitates a quick and easy access to the platform and its complex analysis functions. Input forms are simple and user-friendly. Workflows can be easily customized to specific needs. Experienced users can create their own workflows.
Fully integrated upstream analysis
The platform provides a fully integrated upstream analysis, which combines state-of-the-art analysis of regulatory genome regions with sophisticated pathway analyses.
Knowledge-based data analysis
The platform uses a number of renowned high-quality databases for the data analysis. TRANSFAC® and TRANSPATH® are expert-curated databases. GeneWays is generated by an NLP-based text-mining approach, providing a helpful complement for manually curated data. Well-known public-domain databases like Reactome and HumanCyc are integrated and applied as well.
JavaScript and R scripts
User-specific scripts in JavaScript and in R can be added directly into the platform, and immediately executed. They can be combined with pre-existing analyses, and can be part of the workflows.
NGS data analysis
NGS data analysis is supported by the platform. ChIP-seq data sets containing in vivo transcription factor binding sites or methylation results can be analyzed with the help of ready-made workflows. Galaxy tools are integrated, supporting RNA-seq data analysis, and many functions more.
Simulation engine inside
The platform contains a simulation engine that executes differential equation systems and visualizes the results. Parameter optimization, parameter fitting (based on expression data), and hierarchical modeling are supported.
Videos
Examples
Any user of the geneXplain platform can view the free examples demonstrating the platform abilities towards processing various types of multi-omics data in different studied biological processes and pathologies.
The Examples are located in the Data tab of the geneXplain platform interface under the Examples folder:
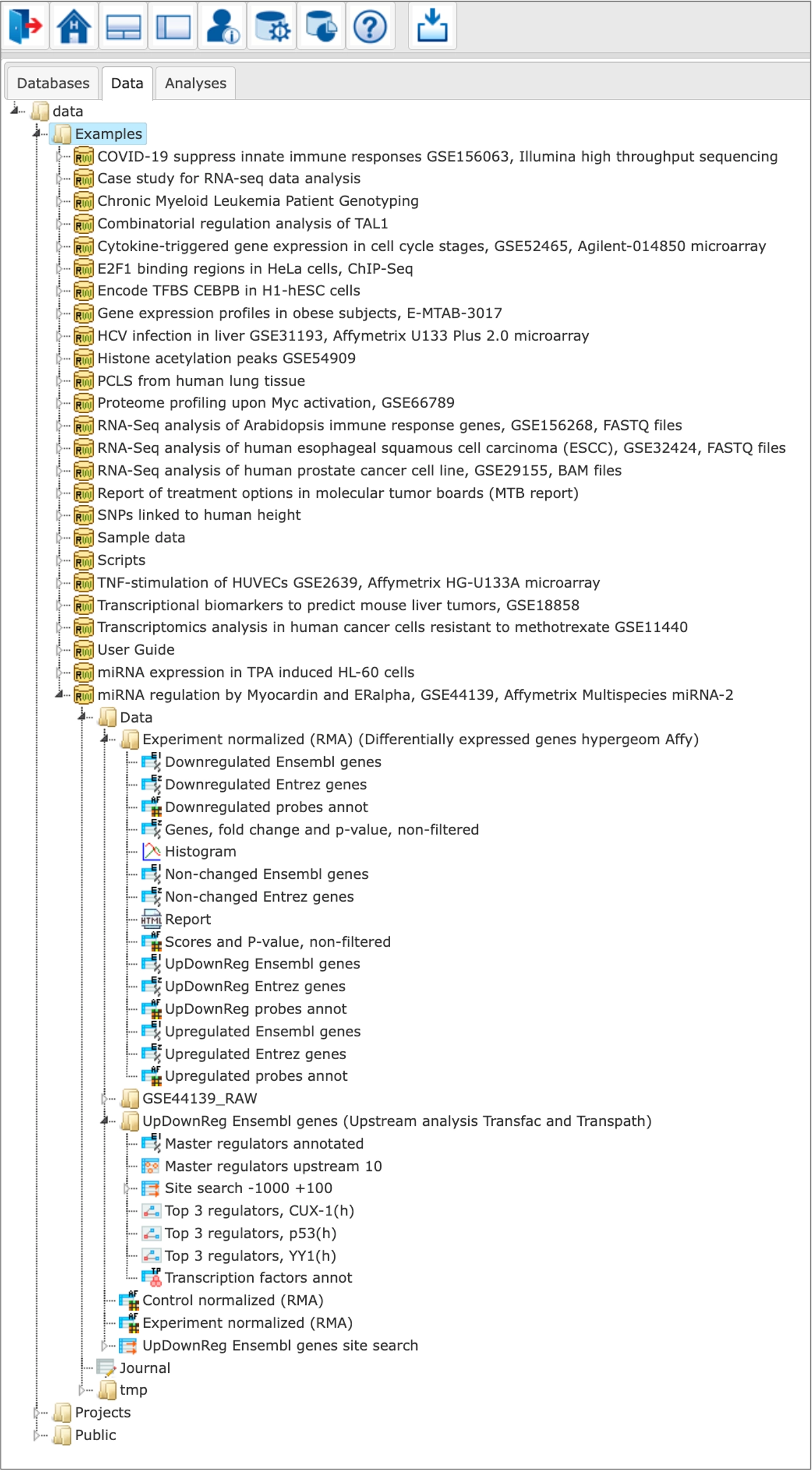
Description of each example is available in the info box upon the click on the name of the respective project:
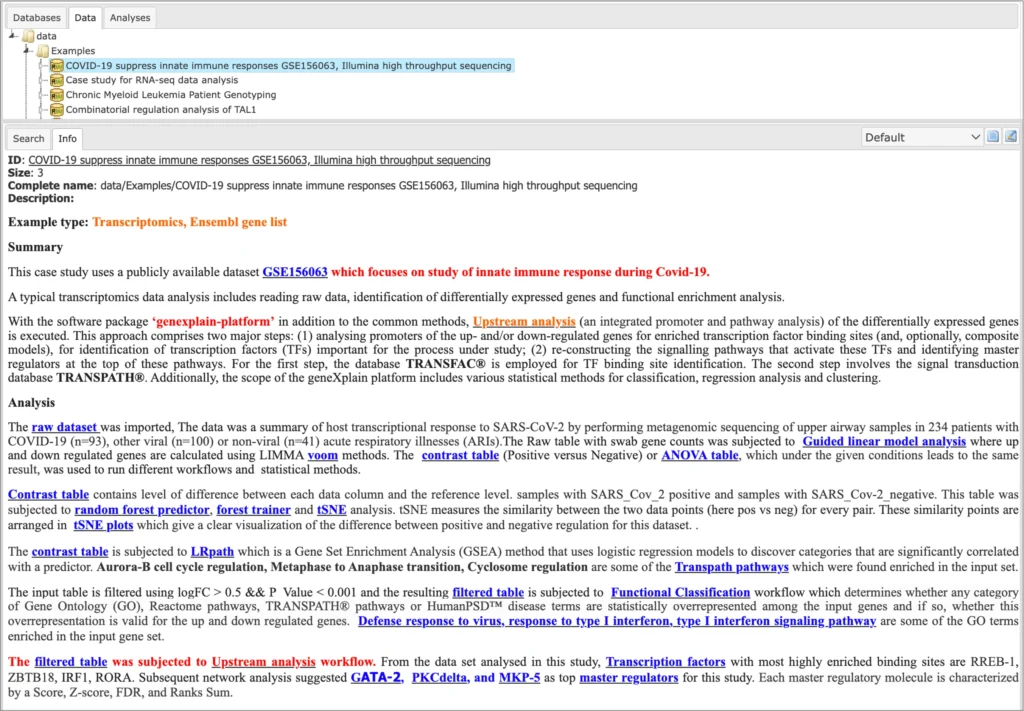
Species Supported:
The geneXplain platform supports analysis across multiple organism groups, including Vertebrata, Plantae, Fungi, Insecta, Nematoda, and Urochordata. Additionally, users can import custom genomes of their species of interest and perform analyses seamlessly.
Methods and workflows for TFBS prediction
The geneXplain platform provides TFBS prediction tools from single sequence to whole genome. In the geneXplain platform interface, you will find a great variety of methods and workflows for TFBS prediction. Below is a structured overview of key workflows, categorized by input type.
General info
The TFBS prediction tools are located in the Analyses –> Methods –> Site analysis section of the platform, as well as in the Analyses –> Workflows –> TRANSFAC section of the platform. Selected tools for sequence analysis can be found under the Sequence analysis button of the platform start page:
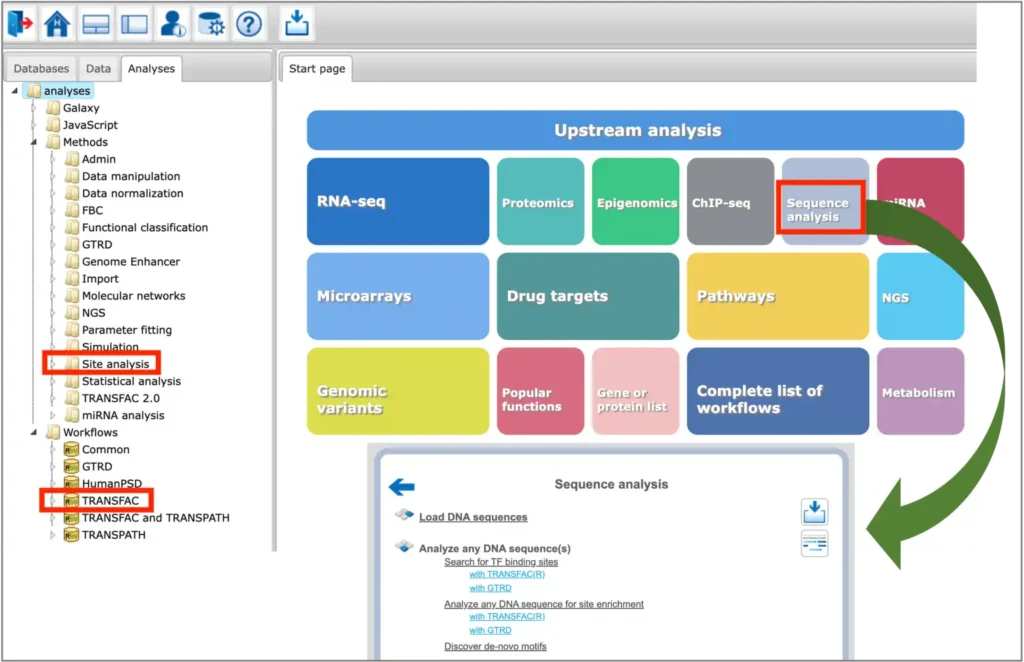
You can find information on the profile selection (collection of positional weight matrices – PWMs – transcription factor binding models that will be used for performing the site search) in this document or in tabular format here.
Sequence-Based TFBS Analysis Methods
Search for TF Binding Sites in DNA Sequences (TRANSFAC®)
The Search for TF binding sites with TRANSFAC® workflow in the geneXplain platform is designed to search for putative transcription factor binding sites, TFBS, in any input DNA sequence in EMBL, Fasta or Genbank formats.
Using this workflow you can analyze DNA sequences of any species and of any genomic regions. In the analysis results of this workflow you will find a summary table and a track with found sites in the input sequences.
Input: DNA sequences (FASTA, EMBL, GenBank)
What you get: – TFBS density per sequence – Detailed binding site tables with positions and scores – Genome browser visualization of TFBS
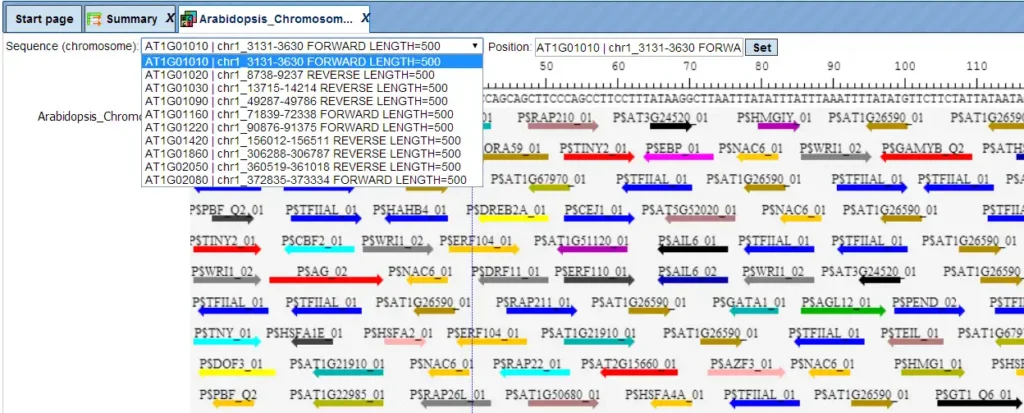
The same track can be opened in a tabular form also.
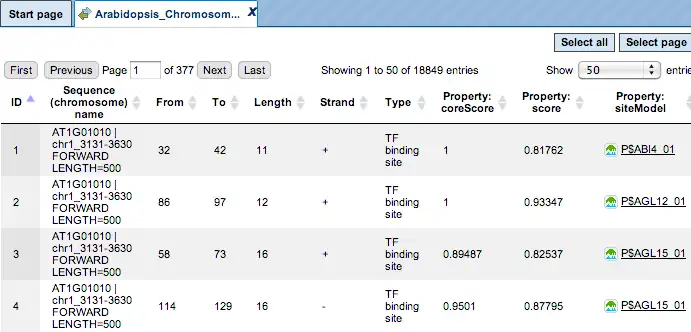
Each row of such table corresponds to one resulting TFBS and includes sequence names, site positions calculated by the algorithm and a site model (TRANSFAC® matrix). This table can be exported as a track in several different formats including intervals, bed, wig and more. DNA sequences can be exported in multi-FASTA format.
Additional visualisation options are available for selected rows of the Summary table: the Report on selected matrices button at the top menu panel of the platform will visualize the found TFBS in the input sequences. In this example, all matrices with a site density <5 were selected. The visualization results are shown below:
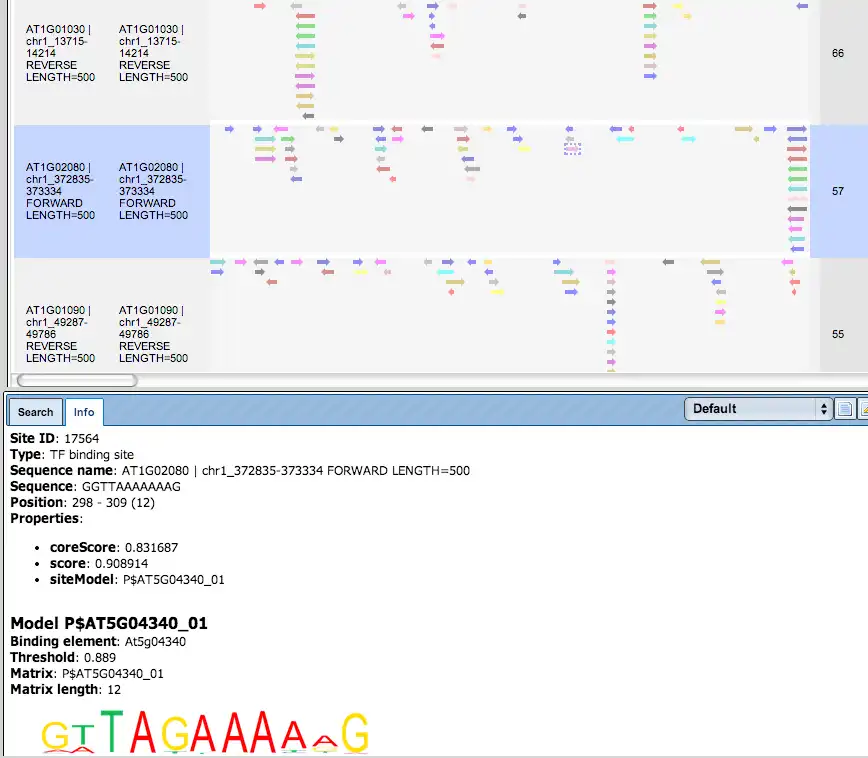
There are ten rows corresponding to the individual sequences in the input set. The column Sites view schematically represents the sequence length with mapped TFBSs. Matches for different matrices are shown in different colors. You can select individual matches by mouse click and get additional information in the Info box.
TFBS Enrichment Analysis (Yes/No Sequence Comparison)
Analyze any DNA sequence for site enrichment with TRANSFAC®
The Analyze any DNA sequence for site enrichment with TRANSFAC® workflow in the geneXplain platform is designed to search for enriched TFBS in any input DNA sequence as compared to a background DNA sequence.
The central part of this workflow is performed by two individual methods: Site search on track and Site search result optimization, both can be found in Analyses –> Methods –> Site analysis.
Input: Target (Yes) sequences vs background (No) sequences of any species. Sequences can be in EMBL, FASTA or GenBank format. What you get: – Enriched TFBS with statistical significance (p-values) – Yes/No ratio for condition-specific binding – Optimized TFBS predictions and visualization tracks
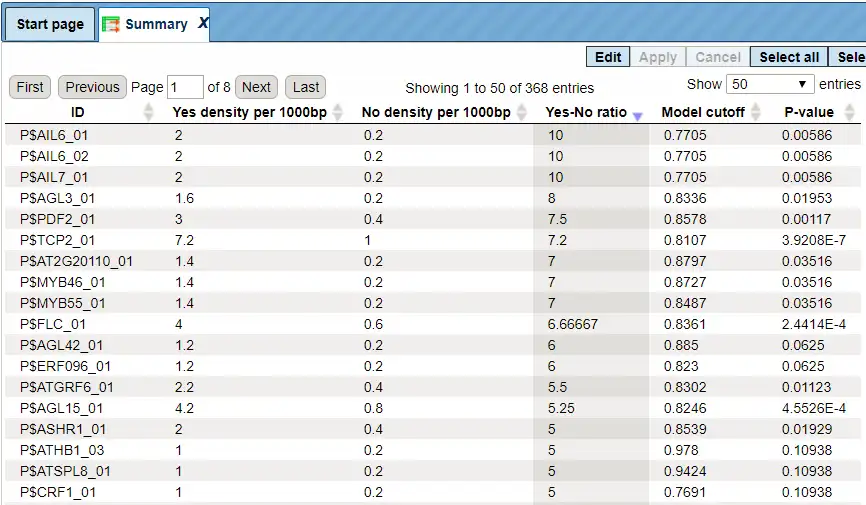
Each row summarizes the information for one site model (PWM – positional weight matrix).
For each row, the columns Yes density per 1000bp and No density per 1000bp show the number of matches normalized per 1000 bp length for the sequences in the input Yes set and input No set, respectively. The Column Yes-No ratio is the ratio of the first two columns. The higher the Yes-No ratio, the higher is the enrichment of matches for the respective matrix in the Yes set. The matrix cutoff values as they are calculated by the program at the optimization step are shown in the column Model cutoff, and the last column shows the p-value of the corresponding event.
TFBSs can be further visualized in the Yes sequences by selecting one or several rows of the Summary table and clicking on the Report on selected matrices button at the top menu panel of the platform.
The track of found sites represents TFBSs that are over-represented in the Yes sequences versus the No sequences. It can be viewed in the genome browser:
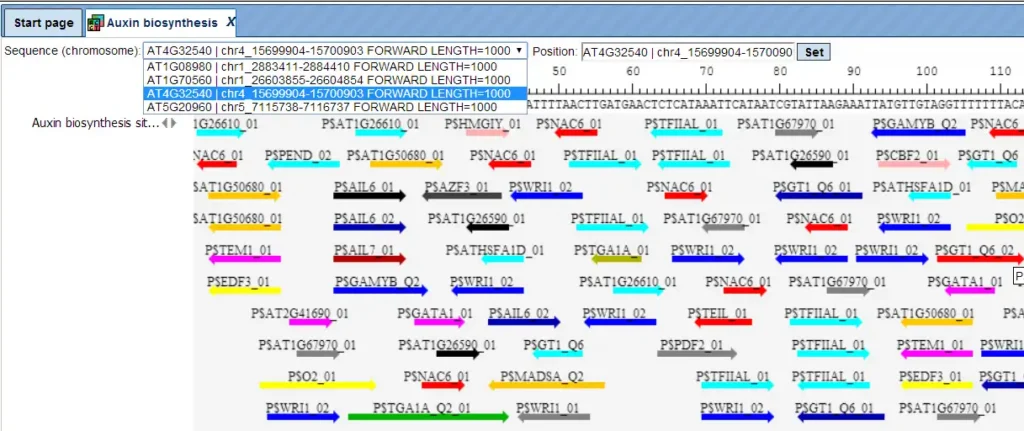
In case of analysis Human, Mouse or Rat data, as well as since recent release the Arabidopsis, Zebrafish, Nematoda, Fruit fly, Baker’s yeast, and Fission yeast data, additional tables will be outputted by the workflow: Transcription factors Ensembl and Transcription factors Entrez. These tables aim at showing transcription factors linked to the identified site models (matrices). These are potential candidate regulators of genes in the input Yes set. They are supposed to regulate transcription of Yes-genes via the identified enriched TFBSs.
You will find a much more detailed description of the sequence analysis workflows in the geneXplain platform in the respective chapter of the user manual.
Gene Set-Based TFBS Analysis
- Site search on gene set
The Site search on gene set method of the geneXplain platform provides you with an ability to search for putative TFBS in a set of genes.
As input for the analysis two gene sets should be provided: Yes (e.g. differentially expressed in an experiment, test set) and No (set of background genes, control set) as well as positional range relative to the TSS and a collection of predefined weight matrices with a particular threshold (profile).
The analysis can be done for Human, Mouse, Rat, Arabidopsis, Nematoda, Zebrafish, Fruit fly, Baker’s yeast, and Fission yeast genes.
Input: Gene sets (Yes vs No), promoter regions relative to TSS
What you get: – Enriched TFBS in promoters of target genes – Identification of candidate transcription factors – Promoter and TFBS visualization tracks
An example of the summary table is shown below:
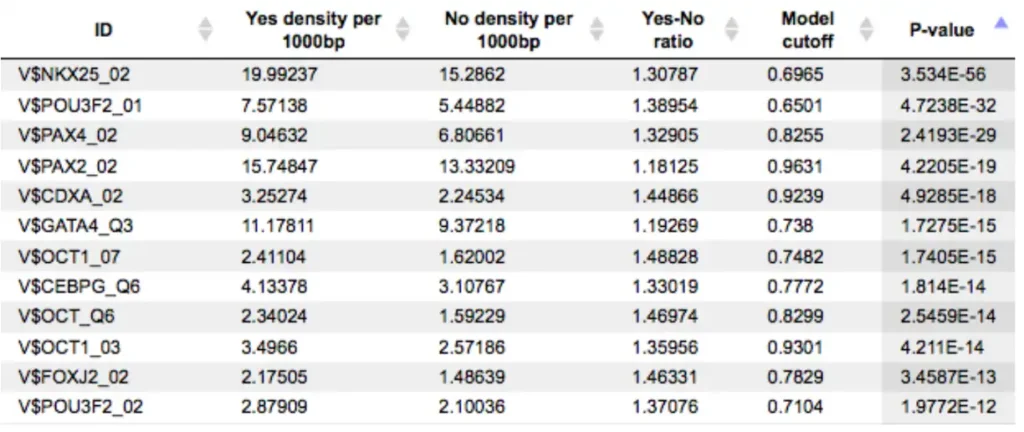
Each row represents one PWM. Columns show TFBS density in Yes and No sets (per 1000 bp), their ratio (Yes/No), and statistical significance. Only matrices with a Yes/No ratio >1 are included, indicating enrichment in the Yes set. Additional columns provide optimized matrix cutoffs and p-values.
Some more snap-shots of the output generated using site search on gene-set.
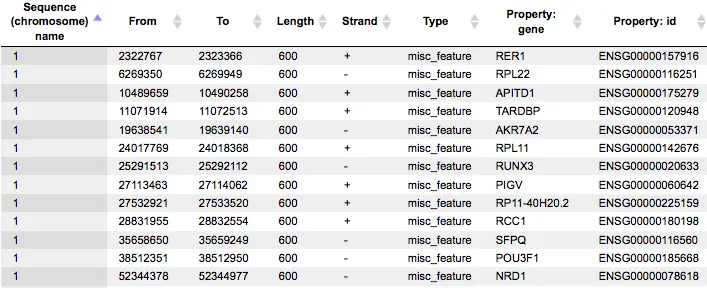
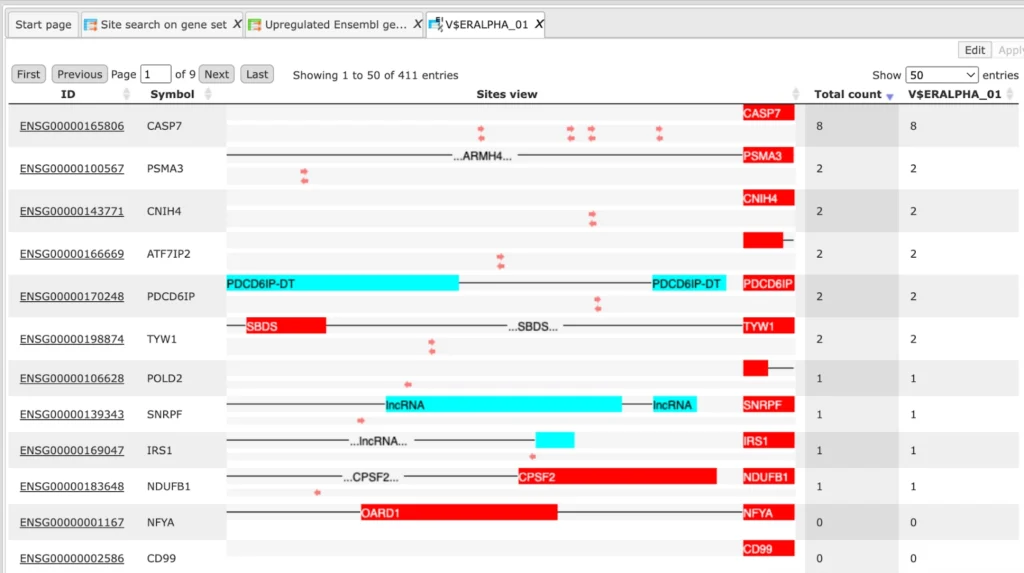
A much more detailed description of the Site search on gene set method can be found in the respective chapter of the geneXplain platform user guide.
Genomic Region / Track-Based TFBS Analysis
Site search on track
The Site search on track method of the geneXplain platform provides you with an ability to search for putative TFBS in an input track.
Input: Genomic coordinates (BED or track format)
What you get: – TFBS mapped to genomic regions – Detailed tables with position, strand, and scores – Exportable tracks for downstream analysis
The result of this method is the track of found sites, which can be visualized as a table:
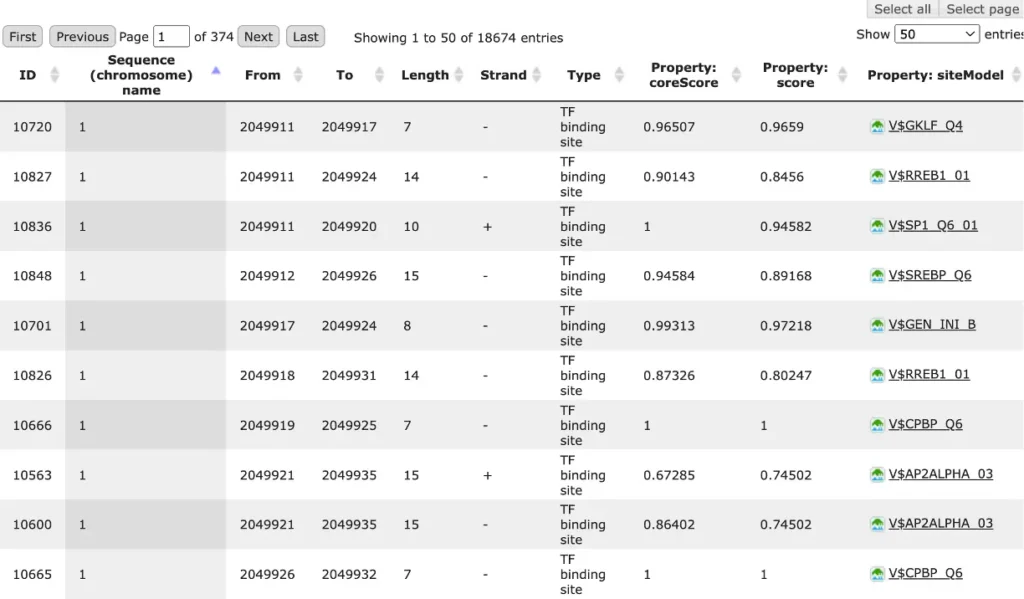
Each row of the table presents details for each individual match for every PWM.
The columns Sequence (chromosome) name, From, To, Length and Strand show, correspondingly, genomic location of the match including chromosome number, start and end positions, strand and length of the match.
The column Type contains information about the type of the elements, in this case all matches are considered as “TF binding site”.
Further columns keep information about PWM producing each match (column Property: siteModel) as well as a score of the core (column Property:coreScore) and a score for the whole matrix (column Property: score).For details about these scores, please see Kel, Alexander E., et al. “MATCH: a tool for searching transcription factor binding sites in DNA sequences.” Nucleic acids research 31.13 (2003): 3576-3579, LINK.
ChIP-seq Based TFBS Analysis Workflows
The geneXplain platform also provides tools for mapping of TFBS to the peaks calculated from ChIP-seq data. You will find respective workflows under the ChIP-seq button of the geneXplain platform main menu:
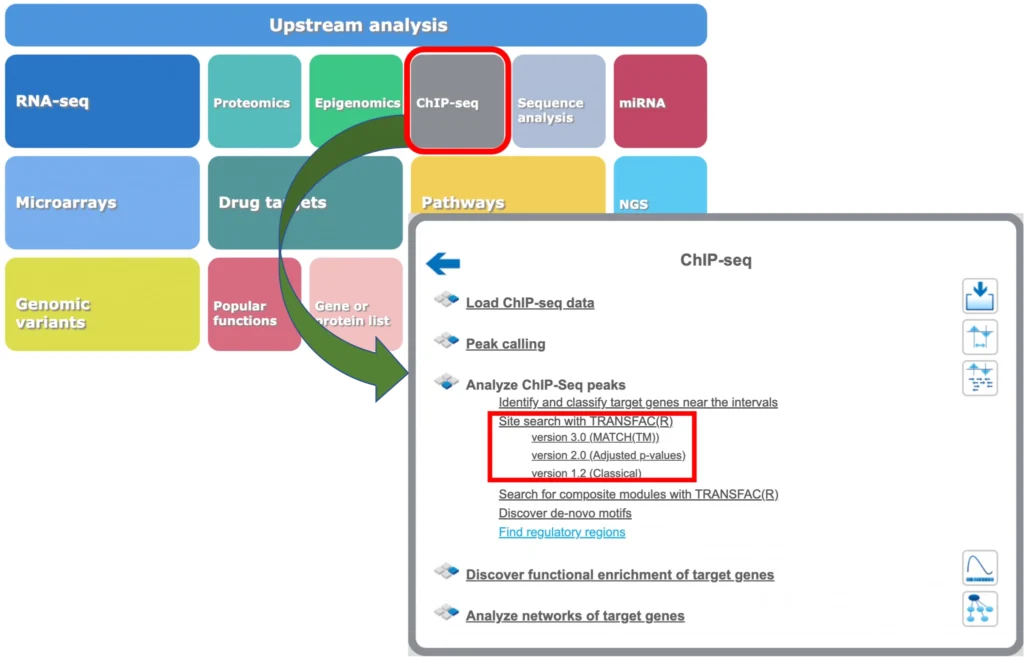
Site search in ChIP-seq peaks: Version 1.2 (Classical)
This workflow helps to map TFBS on peaks calculated from ChIP-seq data.
Site search is done with the help of the TRANSFAC® library of positional weight matrices, PWMs, using the pre-computed profile vertebrate_non_redundant_minSUM.
The input track should be provided in the BED format and submitted to the analysis as the Yes track. The No track can be selected from the ready tracks of housekeeping genes or uploaded as a custom track of your choice.
The results of the workflow contain two tables: Site optimization summary and Transcription factors and two tracks: Yes sites opt and No sites opt.The Site optimization summary table includes the matrices the hits of which are over-represented in the Yes track versus the No track. Only the matrices with Yes-No ratio higher than 1 are included in this output table. The hits of these matrices can be interpreted as over-represented in the Yes set versus No set:
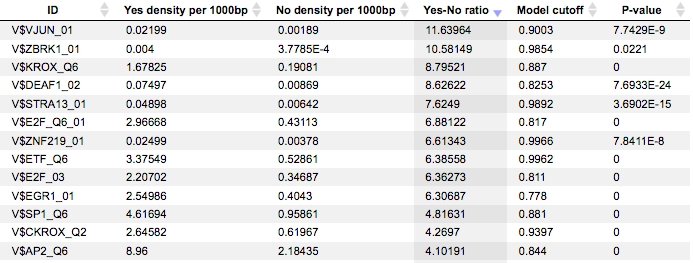
A much more detailed description of this workflow can be found here.
Machine Learning-Based TFBS Discovery (MEALR)
Site search in ChIP-seq peaks: Version 2.0 (Adjusted p-values)
This workflow is designed to map putative enriched TFBSs on peaks calculated from your ChIP-seq data (Yes set) as compared to a random background set (No set). Importantly, the No set is created automatically and contains by default 1000 intervals.
Input: ChIP-seq peaks
What you get: – Enriched motifs with controlled false discovery rate – Optimized TFBS profiles – High-confidence regulatory signals
The workflow first identifies enriched motifs using the MEALR approach. These motifs are used to generate a custom PWM profile, which is then applied to the same ChIP-seq peaks (input as a BED Yes track) to detect enriched TFBS.
Users can select a TRANSFAC® or custom profile (default: vertebrate_non_redundant_minSUM) and apply a coefficient filter (e.g., >0.125 for ~75% TDR) to retain high-confidence motifs.
Outputs include:
- Enriched motifs (MEALR) with coefficient-based ranking
- Generated PWM profile (input-specific)
- TFBS site search summary
- Transcription factors associated with identified motifs
- Tracks for Yes sites and background comparison
Higher MEALR coefficients indicate stronger discriminatory motifs between Yes and No sets.A much more detailed description of this workflow can be found here.
TFBS Enrichment with MATCH™
Site search in ChIP-seq peaks: Version 3.0 (MATCH (TM))
This workflow identifies enriched transcription factor binding sites (TFBSs) in genomic sequences by comparing them to a random background set. It supports genomes such as human, mouse, rat, Arabidopsis, and zebrafish.
Using TRANSFAC® Match™ for tracks method, it identifies
enriched binding sites within the sequences, and outputs a table of potential transcription factors associated with these TFBSs.
Input: Genomic regions and background sequences
What you get: – Enriched TFBS using curated TRANSFAC® matrices – Associated transcription factors – Interpretable tables and genome browser tracks
Genomic sequences in track format (input example) can be submitted for the input. A random track of 1000 sequences that does not overlap with the input sequences is automatically generated as the background set.
The results of the workflow contain several tables and tracks. The identified enriched transcription factor binding sites (TFBSs) are present in a summary table (result example) and can be visualized in the genome browser as a track (result example). The potential transcription factors are given in a final Ensembl table (result example) with annotated GeneSymbol IDs and a short description.
A much more detailed description of this workflow can be found here.
geneXplain platform API
The geneXplain platform provides a comprehensive environment to analyze biomedical and biological data. Its functionality includes, among other things, data storage, data management, data sharing, in-built bioinformatics and systems biology analysis tools, ability to build and run analysis pipelines and workflows, build and visualize molecular network models, or develop quantitative models and perform simulations. More details about the platform can be found on the platform product page and in the corresponding research articles.
The geneXplain platform can be used through a graphical web interface or through APIs (Application Programming Interfaces) that have been implemented in the languages Java and R. Besides being integrated as a software library, the genexplain-api package provides several commandline utilities. The exec tool allows to configure and run remote analyses using JSON input files. The JSON interface is described in detail in the genexplain-api documentation.
The following figure sketches some of the tasks that can be carried out with the platform APIs:
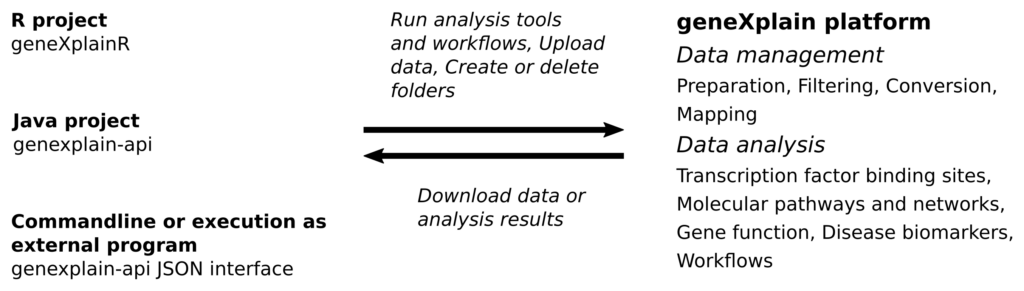
All analysis and utility tools provided by the geneXplain platform as well as integrated Galaxy tools and workflows can be executed using API functions. In addition, there are methods to manage, organize and download research data and analysis results. Notably, analysis jobs can run asynchronously so that you are not required to wait for long analysis tasks.
You are welcome to view the geneXplain platform API tutorial covering both: the geneXplain platform R and the geneXplain platform Java APIs, as well as the command line interface, here.
geneXplain platform R API (geneXplainR)
GeneXplainR package provides an interface for the easier integration of the geneXplain platform functionalities into the R pipelines. Combine the wide range of data, tools, and workflows of the geneXplain platform with the resources available in R language to perform your bioinformatic analysis.
The geneXplainR package provides an R client for the geneXplain platform, geneXplainR is based on and extends the rbiouml package. A goal of this project is to add functionality that helps to make building R pipelines that use the geneXplain platform easier.
Watch this video demonstrating the example of executing the “Identify enriched motif in promoters (TRANSFAC)” workflow of the geneXplain platform using the R API:
Check out the record of Coffee break with TRANSFAC webinar devoted to the geneXplain platform API:
geneXplain platform Java API
The geneXplain Java API allows integration of geneXplain platform functionalities into Java programs and provides a JSON interface to configure and execute individual analysis tasks as well as complex workflows. The geneXplain platform Java API can be used to write Java programs that can invoke functionality of the platform through its web interface, e.g. for import and export of biological data, or to submit and monitor analysis jobs. Furthermore, platform tasks can be specified in JSON format and submitted using the executable JAR file. Though not endowed with all possibilities of a programming language, the JSON interface facilitates, among other things, definition of templates, branch points, or nesting of tasks, so that you can build complex workflows from reusable components. It is intended to be applied as part of a dynamic and polyglot analysis environment that utilizes diverse programming languages and resources.
Watch thIs introductory tutorial on geneXplain platform Java API that shows how to install the API software and run the basic commands:
Machine Learning and AI methods
TRANSFAC integrates advanced tools for the prediction of transcription factor binding sites (TFBS) and their combinatorial patterns across the genome using machine learning and AI-based approaches.
These capabilities include:
- AI-driven combinatorial TFBS analysis using the Composite Module Analyst algorithm
- Combinatorial modeling based on sparse logistic regression (MEALR)
AI-driven combinatorial TFBS analysis using Composite Module Analyst algorithm
Introduction to Composite Modules
Composite modules represent combinations of multiple TFBS that co-occur within regulatory DNA sequences. Identifying these modules enables the discovery of regulatory patterns that are significantly overrepresented in a target dataset compared to a background set.
Within the geneXplain platform, composite modules are identified using an in-house implementation of a genetic algorithm known as the Composite Module Analyst
Composite Module Analysis Workflow
The Composite Module Analyst operates on the output generated from TFBS site search analyses. Two complementary analysis workflows are available:
1. Construct Composite Modules (Gene Set-Based)
This method analyzes promoter sequences defined relative to transcription start sites (TSS) for a given gene set.
- Input: Results from Site search on gene set analysis
- Application: Identification of TFBS combinations in promoter regions
- Output: Composite modules that distinguish a target gene set (Yes-set) from a background set (No-set)
2. Construct Composite Modules on Tracks (Genome Coordinate-Based)
This method operates on DNA sequences defined by absolute genomic coordinates and is particularly suited for high-throughput sequencing data.
- Input: Results from Site search on track analysis
- Typical use case: Analysis of ChIP-seq fragments
- Output: Composite modules distinguishing a target track (Yes-track) from a background track (No-track)
Key Differences Between the Methods
The two approaches differ primarily in:
- Type of input sequences
- Gene-based promoter regions vs. genomic coordinate-defined tracks
- Input data format
- Gene sets vs. genomic tracks
Access in geneXplain Platform :
Both analysis workflows are available in:
Analyses → Methods → Site analysis
Summary of Composite Module Construction
- The gene set-based method identifies regulatory TFBS combinations in promoter regions, distinguishing between Yes-set and No-set gene groups.
- The track-based method identifies TFBS combinations in genomic regions such as ChIP-seq data, distinguishing between Yes-track and No-track datasets.
Both analyses require:
- Precomputed site search results
- Defined Yes/No datasets
- A selected matrix profile
Additional Information
Hierarchical Structure of Composite Modules
Before interpreting the results, it is important to understand how composite modules are structured and visualized.
Composite modules are organized in a two-level hierarchy:
- Site models
- Modules
At the highest level, multiple modules together form a promoter model, representing the regulatory architecture of a gene.
Site Models (Level 1)
Site models represent individual transcription factor binding models derived from PWMs used in the selected profile. Each site model includes:
- A threshold value optimized by the genetic algorithm
- An N value representing the number of best TFBS matches used for scoring
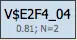
Modules (Level 2)
Modules group multiple site models into combinatorial regulatory units. Each module:
- Contains several site models
- Is defined by a module width (DNA region where TFBS cluster)
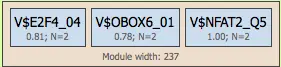
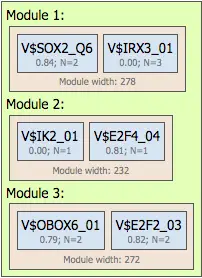
Promoter Model
Multiple modules form a promoter model describing gene regulation. Users can define:
- Number of modules
- Number of site models per module
- Number of TFBS considered
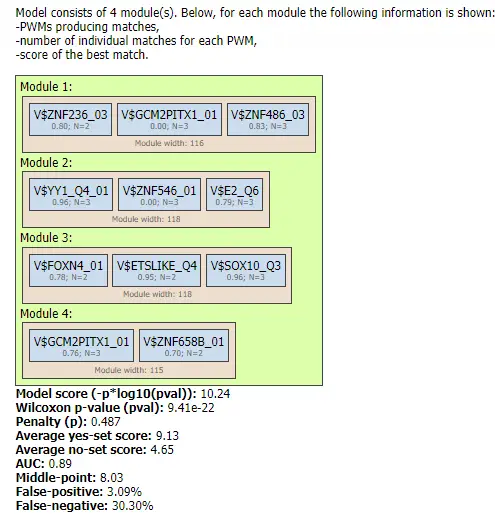
Results Visualization
Below we provide the results visualisation of the Construct composite modules analysis obtained for the demo input data set. The input parameters of the method that were used for the analysis launch were as follows:

As a result, the method constructed two tables (Model visualization on Yes set and Model visualization on No set), two tracks (Yes track and No track), and one histogram.
In the Model visualization on Yes set table the the primary results of the analysis are presented: the identified composite modules are shown in the promoters of the Yes set:

Each row in this table corresponds to one gene of the Yes set, and for each gene the Ensembl ID and the gene symbol are shown in the two first columns. The column Model displays a symbolic map of the gene promoter taken for the analysis, in this case -500/+100 relative to the TSS.
Arrows of different colors correspond to individual TFBSs, and a gradient in grey corresponds to the statistical density of the identified composite modules. The most intensive grey color corresponds to the center of a composite module. Each individual TFBS on this map is clickable, and upon a click information is displayed in the Info box (bottom left corner in the tool).
As an example, one blue arrow is selected on the promoter of the top gene in the screenshot above, and for this selected TFBS the following details are shown in Info box:

The last column in the table, Score, shows a score calculated for each promoter depending on the number of modules, site models, sites, their scores and other statistical parameters. The higher the score for a promoter, the better the differentiation of this promoter from the promoters of the No set. The column Score is used for default sorting of the table, with the highest scores on top.
In addition to that, at the bottom part of the tool in the Model visualization on Yes set table you can also see the schematic representation of the hierarchical structure of the identified composite module, as well as a comprehensive set of its statistical characteristics:

The Yes track provides essential information about the regulation of individual promoters and is therefore important to be included in the visualization of individual promoters by the genome browser.
The schematic visualization can be comfortably extended to a more detailed visualization for each individual promoter:
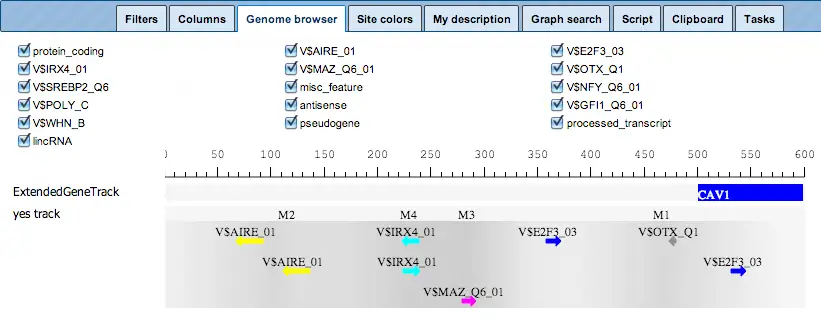

For a selected promoter, you can see a more detailed map, including the names of the matrices and the numbers of individual modules, M1 through M4. Each element of this interactive map has a corresponding check box. Unchecked elements will not be displayed on the map. De-selection is applied simultaneously to both: the detailed view of one promoter, and the table with the schematic representation of all promoters.
The table Model visualization on No set shows a visualization of the identified composite modules in the promoters of the No set.
The structure of this table is the same as that of the Model visualization on Yes set table, described above.
The function of the No track is to provide a possibility for a detailed visualization of no promoters in a way similar to that of the Yes track.
The distribution of scores for individual promoters is shown as a Histogram, where the promoter score value is shown on X axis and the percentage of promoters (% sequences) having this score is shown on the Y axis:
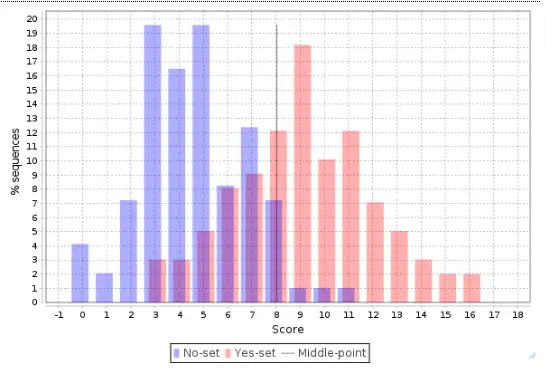
This histogram can be further interpreted applying the statistical characteristics described above.
The center, a vertical grey line, corresponds to the average score value and is equal to 3.44 in this example. Promoters from the No set with a score above 3.44 are shown in the histogram as blue bars to the right of the center, and they are referred to as false positives. In this example, the false positive rate is 16.82 %.
Promoters from the Yes set with a score below 3.44 are shown in the histogram as red bars to the left of the center, and they are referred to as false negatives. In this example, the false negative rate is 23.42 %.
A visual analysis of the histogram suggests that the Yes promoters with a score above 4.5 are very well separated from the No promoters, which means that for this part of the promoters the composite model constructed is most suitable. In this example there are 38 promoters with the score value >4.5; they can be saved as a separate gene set, and for them the model obtained works best.
Score calculation of the composite models
The figure below demonstrates the calculation of the score value for the composite modules in the promoter sequences. The TSS is shown as a thin arrow on the right side of the figure. Four thick arrows exemplify four sites found in this promoter. The color of the arrows exemplifies the site model which these sites belong to (three site models – red, green and blue).
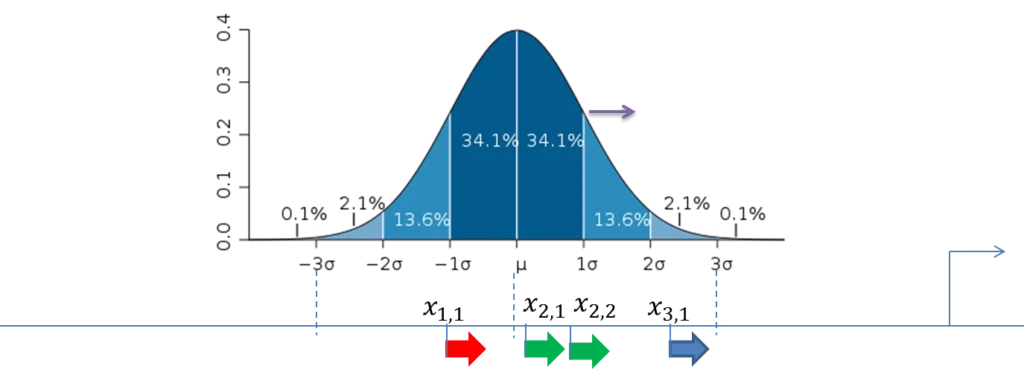
A promoter model consists of K modules. The score of each module Mk(Score(Mk),k= 1, …,K) is calculated according to this formula:

Here, Site Score (t,i) is the site score for the sites found in the promoter, which is calculated by the Match algorithm.
mt– the number of sites of the site model t found in the promoter.
Tk– the number of site models in the module Mk, and
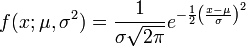
The final promoter score is calculated as the sum of the module scores Mk.
Standard deviation (σ) of the normal distribution is subject of optimization by the genetic algorithm and represents the width of the module in the output of the composite module analysis.
Further details on the composite modules construction can be found in the respective chapter of the geneXplain platform user guide.
Combinatorial modeling based on sparse logistic regression (MEALR)
Combinatorial regulation analysis of genomic or custom sequences
This workflow analyzes DNA sequences to identify combinations of transcription factor (TF) binding regions using MEALR models from TRANSFAC®.
Best suited for:
- Single sequence analysis
- Identifying regulatory patterns in genomic regions
Workflow steps:
The workflow proceeds through the following main steps:
- Prediction of potential binding locations using the TRANSFAC® MEALR combinatorial regulation analysis
- Extraction of TRANSFAC® PWMs represented by MEALR model hits using the Extract TRANSFAC® PWMs from combinatorial regulation analysis
- Preparation of a cutoff profile with extracted PWMs for subsequent MATCHTM search using the Create profile from site model table
- Prediction of binding sites represented by PWMs in input sequences using the TRANSFAC® MATCHTM for tracks
- The tools Filter track by condition and Intersect tracks are applied to derive filtered model predictions and intersections of predicted TF binding site and combinatorial model locations.
Challenges:
Predicting TF binding sites is challenging because:
- Binding sequences are very short (10–20 base pairs)
- They vary significantly in sequence patterns
- Binding depends on biological context (cell type, chromatin state, TF cooperation)
Key advantage:
With almost 1000 human TFs, over 300 cell types and more than 50 tissue types, the TRANSFAC® library of MEALR models provides the first comprehensive collection of TF binding models that account for combinatorial TF-DNA complexes comprising multiple DNA-binding specificities as well as cellular and tissue-related contexts.
These models can therefore deliver predictions with increased accuracy, so that subsequent searches for binding sites that mark the locations of individual TFs are able to focus on relevant PWMs.
This is an important step to select from the large number of PWMs curated by TRANSFAC® and to prioritize binding sites of interest. Within longer sequences the MEALR model predictions furthermore suggest subregions where binding by certain TFs most likely occurs.
A much more detailed description of this workflow can be found here.
MEALR combinatorial regulation analysis
This analysis applies combinatorial regulatory models (CRMs) based on the MEALR affinity score [1] to classify or scan sequences for occurrences of combinations of transcription factor binding sites represented by TRANSFAC® PWMs. The models are taken from the MEALR library whose training data originate from the TRANSFAC® collection of high-throughput sequencing experiments.
The method can be launched either in a classification or in the scan mode. The Classification mode evaluates input sequences as a whole, whereas the scan mode analyzes sequence windows separated by the given step size (sliding window). In scan mode, the Best hit method reports the best scoring sequence window disregarding a cutoff and the Cutoff method reports the best non-overlapping windows satisfying the specified cutoff.
The output folder encompasses a table and sequence track with information about model hits. The output table contains sequence start and end points of hits, model ids, match probabilities as well as other values as described below. For input sequences derived from genomic regions (instead of imported as custom sequences) the table includes in addition a sequence id generated for a region as well as the genomic sequence id, start and end coordinates.
[1] Katie Lloyd, Stamatia Papoutsopoulou, Emily Smith, Philip Stegmaier, Francois Bergey, et al., The SysmedIBD Consortium; Using systems medicine to identify a therapeutic agent with potential for repurposing in inflammatory bowel disease. Dis Model Mech 1 November 2020; 13 (11): dmm044040.
A much more detailed description of this method can be found here.
Extract TRANSFAC® PWMs from combinatorial regulation analysis
This tool extracts TRANSFAC® PWMs from a result table generated by the MEALR combinatorial regulation analysis. The PWMs represent transcription factor binding specificities that constitute the combinatorial module predicted by the MEALR model.
The output contains the TRANSFAC® PWMs extracted from MEALR models according to specified cutoffs. This table can further be applied in several analyses, e.g. to extract corresponding transcription factors using the tool or to create a profile for binding site predictions ( Create profile from site model table) with MATCHTM.
A much more detailed description of this method can be found here.
Search for discriminative sites with TRANSFAC® (MEALR)
The tool MEALR finds combinations of TFBS matrices that discriminate between two sets of sequences (denoted asYes and No sets). TheYes set may consist of genomic regions identified in a ChIP-seq experiment. No sequences are often other non-coding genomic regions not overlapping with the peaks.
MEALR differs from other tools in the following points:
- No cutoff or threshold is used on matrix scores to determine potential binding sites. Instead, MEALR calculates threshold-free sequence scores.
- MEALR builds a discriminative model for classification which is well-established and widely applied in statistical analysis called Sparse Logistic Regression. The model consists of a linear model that estimates the probability that a sequence belongs to the Yes set based on its binding site features.
- The sparseness constraint enables MEALR to select a subset of matrices relevant for classification of Yes and No sequences from a possibly large matrix library. Therefore MEALR’s output differs from other tools by presenting a focused set of matrices.
- While other site enrichment tools provided in the platform evaluate enrichment separately for each matrix, the model used in MEALR assesses the importance of matrices for discrimination in combination with other matrices of the library. Therefore, MEALR suggests (linear) combinations of transcription factor motifs.
For each input sequence x MEALR calculates a score for each PWM by the following equation where W denotes the number
of windows scored by the PWM and LSmax(xw) is the high log-odds score of window w :
f(x) = log( Σw exp(LSmax(xw)) / W )
Each sequence is therefore associated with a vector of scores, one from each matrix, and a class (Yes, No).
Let us present an example analysis for a ChIP-seq data set consisting of 500 peak regions and 1000 sequences randomly sampled from regulatory regions across the human genome.
Yes set is the set of sequence intervals that you want to analyze, for example these can be ChIP-seq peak regions.
No set is the set of background intervals (control set).
In the input profile (collection of PWMs – binding motifs) the cutoffs will be ignored, because MEALR calculates whole sequence scores.
In the results of analysis the summary table would look like this:
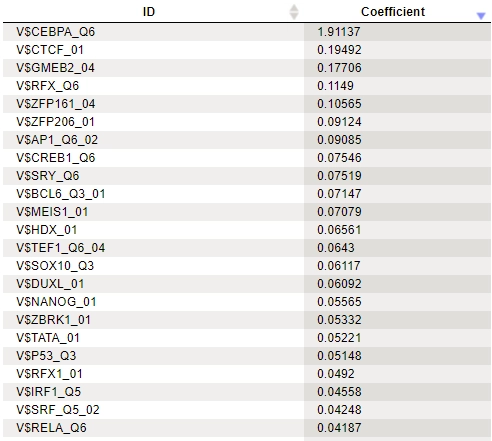
A row of this table contains a matrix identifier and its logistic regression coefficient. The larger the coefficient value, the more important the corresponding matrix was for discriminating between Yes and No sequences. In our example, three of the five top matrices represent members of the transcription factor subfamily C/EBP.
For further details on this method please refer to the respective chapter of the geneXplain platform user guide.
geneXplain platform Applications in Research — Selected Publications
Novikova S., Tolstova T., Kurbatov L., Farafonova T., Tikhonova O., Soloveva N., Rusanov A., Zgoda V. (2024) Systems Biology for Drug Target Discovery in Acute Myeloid Leukemia. Int. J. Mol. Sci. 25(9), 4618 Link
Kisakol, B., Matveeva, A., Salvucci, M., Kel, A., McDonough, E., Ginty, F., Longley, D., Prehn, J. (2024) Identification of unique rectal cancer-specific subtypes. Br J Cancer. 130, 1809–1818. DOI https://doi.org/10.1038/s41416-024-02656-0. Link
Xinxin Liu., Zhihua Huang., Qiuzheng Chen., Kai Chen., Weikang Liu., Guangnian Liu., Xiangyu Chu., Dongqi Li., Yongsu Ma., Xiaodong Tian., Yinmo Yang. (2024) Hypoxia-induced epigenetic regulation of miR-485-3p promotes stemness and chemoresistance in pancreatic ductal adenocarcinoma via SLC7A11-mediated ferroptosis. Cell Death Discovery. 10, 262. Link
Drake, C., Zobl W., Wehr M., Koschmann J., De Luca D., Kühne B. A. , Vrieling H. , Boei J. , Hansen T. , Escher S. E. (2023) Substantiate a read-across hypothesis by using transcriptome data—A case study on volatile diketones. Front. Toxicol. 5. Link
Rajavel A., Klees S., Hui Y., Schmitt A.O., Gültas M. (2022) Deciphering the Molecular Mechanism Underlying African Animal Trypanosomiasis by Means of the 1000 Bull Genomes Project Genomic Dataset. Biology (Basel). 11(5), 742. Link
Menck K., Wlochowitz D., Wachter A., Conradi L.C., Wolff A., Scheel A.H., Korf U., Wiemann S., Schildhaus H.U., Bohnenberger H., Wingender E., Pukrop T., Homayounfar K., Beißbarth T., Bleckmann A. (2022) High-Throughput Profiling of Colorectal Cancer Liver Metastases Reveals Intra- and Inter-Patient Heterogeneity in the EGFR and WNT Pathways Associated with Clinical Outcome. Cancers 14(9), 2084. Link
Myer P.A., Kim H., Blümel A.M., Finngan E., Kel A., Thompson T.V., Greally J.M., Prehn J.H., O’Connor D.P., Friedman R.A., Floratos A., Das S. (2022) Master Transcription Regulators and Transcription Factors Regulate Immune-Associated Differences Between Patients of African and European Ancestry With Colorectal Cancer. Gastro Hep Adv. 1(3), 328–341. Link
Kawashima Y., Nagai H., Konno R., Ishikawa M., Nakajima D., Sato H., Nakamura R., Furuyashiki T., Ohara O. (2022) Single-Shot 10K Proteome Approach: Over 10,000 Protein Identifications by Data-Independent Acquisition-Based Single-Shot Proteomics with Ion Mobility Spectrometry. J Proteome Res. 21(6), 1418–1427. Link
Klees S., Schlüter J.S., Schellhorn J., Bertram H., Kurzweg A.C., Ramzan F., Schmitt A.O., Gültas M. (2022) Comparative Investigation of Gene Regulatory Processes Underlying Avian Influenza Viruses in Chicken and Duck. Biology (Basel). 11(2), 219. Link
Benjamin, S.J., Hawley, K.L., Vera-Licona, P., La Vake, C.J., Cervantes, J.L., Ruan, Y., Radolf, J.D., Salazar, J.C. (2021) Macrophage mediated recognition and clearance of Borrelia burgdorferi elicits MyD88-dependent and -independent phagosomal signals that contribute to phagocytosis and inflammation. BMC Immunol. 22, 32. Link
Menck K., Heinrichs S., Wlochowitz D., Sitte M., Noeding H., Janshoff A., Treiber H., Ruhwedel T., Schatlo B., von der Brelie C., Wiemann S., Pukrop T., Beißbarth T., Binder C., Bleckmann A. (2021) WNT11/ROR2 signaling is associated with tumor invasion and poor survival in breast cancer. J Exp Clin Cancer Res. 40, 395. Link
Meier, T., Timm, M., Montani, M., Wilkens, L. (2021) Gene networks and transcriptional regulators associated with liver cancer development and progression. BMC Med. Genomics 14, 41. Link
Chereda H., Bleckmann A., Menck K., Perera-Bel J., Stegmaier P., Auer F., Kramer F., Leha A., Beißbarth T. (2021) Explaining decisions of graph convolutional neural networks: patient-specific molecular subnetworks responsible for metastasis prediction in breast cancer. Genome Med. 13, 42. Link
Heinrich F., Ramzan F., Rajavel A., Schmitt A.O., Gültas M. (2021) MIDESP: Mutual Information-Based Detection of Epistatic SNP Pairs for Qualitative and Quantitative Phenotypes. Biology (Basel). 10(9), 921. Link
Tenesaca S., Vasquez M., Alvarez M., Otano I., Fernandez-Sendin M., Di Trani C.A., Ardaiz N., Gomar C., Bella A., Aranda F., Medina-Echeverz J., Melero I., Berraondo P. (2021) Statins act as transient type I interferon inhibitors to enable the antitumor activity of modified vaccinia Ankara viral vectors. J Immunother Cancer. 9(7), e001587. Link
Vanvanhossou S.F.U., Giambra I.J., Yin T., Brügemann K., Dossa L.H., König S. (2021) First DNA Sequencing in Beninese Indigenous Cattle Breeds Captures New Milk Protein Variants. Genes (Basel). 12(11), 1702. Link
Lloyd K., Papoutsopoulou S., Smith E., Stegmaier P., Bergey F., Morris L., Kittner M., England H., Spiller D., White M.H.R., Duckworth C.A., Campbell B.J., Poroikov V., Martins Dos Santos V.A.P., Kel A., Muller W., Pritchard D.M., Probert C., Burkitt M.D.; SysmedIBD Consortium. Using systems medicine to identify a therapeutic agent with potential for repurposing in inflammatory bowel disease. Dis Model Mech. 13(11), dmm044040. Link
Odagiu L., Boulet S., Maurice De Sousa D., Daudelin J.F., Nicolas S., Labrecque N. (2020) Early programming of CD8+ T cell response by the orphan nuclear receptor NR4A3. Proc Natl Acad Sci U S A. 117(39), 24392–24402. Link
Ayyildiz D., Antoniali G., D’Ambrosio C., Mangiapane G., Dalla E., Scaloni A., Tell G., Piazza S. (2020) Architecture of The Human Ape1 Interactome Defines Novel Cancers Signatures. Sci Rep. 10, 28. Link
Ural, B.B., Yeung, S.T., Damani-Yokota, P., Devlin, J.C., de Vries, M., Vera-Licona, P., Samji, T., Sawai, C.M., Jang, G., Perez, O.A., Pham, Q., Maher, L., Loke, P., Dittmann, M., Reizis, B., Khanna, K.M. (2020) Identification of a nerve-associated, lung-resident interstitial macrophage subset with distinct localization and immunoregulatory properties. Sci. Immunol. 5, eaax8756. Link
Leiherer A., Muendlein A., Saely C.H., Fraunberger P., Drexel H. (2019) Serotonin is elevated in risk-genotype carriers of TCF7L2 – rs7903146. Sci Rep. 9, 12863. Link
Wang B., Ran Z., Liu M., Ou Y. (2019) Prognostic Significance of Potential Immune Checkpoint Member HHLA2 in Human Tumors: A Comprehensive Analysis. Front Immunol. 10, 1573. Link
Mekonnen, Y.A., Gültas, M., Effa, K., Hanotte, O., Schmitt, A.O. (2019) Identification of Candidate Signature Genes and Key Regulators Associated With Trypanotolerance in the Sheko Breed. Front. Genet. 10, 1095. Link
Blazquez, R., Wlochowitz, D., Wolff, A., Seitz, S., Wachter, A., Perera-Bel, J., Bleckmann, A., Beißbarth, T., Salinas, G., Riemenschneider, M.J., Proescholdt, M., Evert, M., Utpatel, K., Siam, L., Schatlo, B., Balkenhol, M., Stadelmann, C., Schildhaus, H.U., Korf, U., Reinz, E., Wiemann, S., Vollmer, E., Schulz, M., Ritter, U., Hanisch, U.K., Pukrop, T. (2018) PI3K: A master regulator of brain metastasis-promoting macrophages/microglia. Glia 66, 2438-2455. Link
Orekhov, A.N., Oishi, Y., Nikiforov, N.G., Zhelankin, A.V., Dubrovsky, L., Sobenin, I.A., Kel, A., Stelmashenko, D., Makeev, V.J., Foxx, K., Jin, X., Kruth, H.S. Bukrinsky, M. (2018) Modified LDL Particles Activate Inflammatory Pathways in Monocyte-derived Macrophages: Transcriptome Analysis. Curr. Pharm. Des. 24, 3143-3151. Link
Smetanina, M.A., Kel, A.E., Sevost’ianova, K.S., Maiborodin, I.V., Shevela, A.I., Zolotukhin, I.A., Stegmaier, P., Filipenko, M.L. (2018) DNA methylation and gene expression profiling reveal MFAP5 as a regulatory driver of extracellular matrix remodeling in varicose vein disease. Epigenomics 10, 1103-1119. Link
Kalozoumi, G., Kel-Margoulis, O., Vafiadaki, E., Greenberg, D., Bernard, H., Soreq, H., Depaulis, A., Sanoudou, D. (2018) Glial responses during epileptogenesis in Mus musculus point to potential therapeutic targets. PLoS One 13, e0201742. Link
Mandić, A.D., Bennek, E., Verdier, J., Zhang, K., Roubrocks, S., Davis, R.J., Denecke, B., Gassler, N., Streetz, K., Kel, A., Hornef, M., Cubero, F. J., Trautwein, C. and Sellge, G. (2017) c-Jun N-terminal kinase 2 promotes enterocyte survival and goblet cell differentiation in the inflamed intestine. Mucosal Immunol. 10, 1211-1223. Link
Niehof, M., Hildebrandt, T., Danov, O., Arndt, K., Koschmann, J., Dahlmann, F., Hansen, T. and Sewald, K. (2017) RNA isolation from precision-cut lung slices (PCLS) from different species. BMC Res. Notes 10, 121. Link
Triska, M., Solovyev, V., Baranova, A., Kel, A., Tatarinova, T.V. (2017) Nucleotide patterns aiding in prediction of eukaryotic promoters. PLoS One 12, e0187243. Link
Pietrzyńska, M., Zembrzuska, J., Tomczak, R., Mikołajczyk, J., Rusińska-Roszak, D., Voelkel, A., Buchwald, T., Jampílek, J., Lukáč, M., Devínsky, F. (2016) Experimental and in silico investigations of organic phosphates and phosphonates sorption on polymer-ceramic monolithic materials and hydroxyapatite. Eur. J. Pharm. Sci. 93, 295-303. Link
Ciribilli, Y., Singh, P., Inga, A., Borlak, J. (2016) c-Myc targeted regulators of cell metabolism in a transgenic mouse model of papillary lung adenocarcinoma. Oncotarget 7, 65514-65539. Link
Wlochowitz, D., Haubrock, M., Arackal, J., Bleckmann, A., Wolff, A., Beißbarth, T., Wingender, E., Gültas, M. (2016) Computational Identification of Key Regulators in Two Different Colorectal Cancer Cell Lines. Front. Genet. 7, 42. Link
Lee, E.H., Oh, J.H., Selvaraj, S., Park, S.M., Choi, M.S., Spanel, R., Yoon, S. and Borlak, J. (2016) Immunogenomics reveal molecular circuits of diclofenac induced liver injury in mice. Oncotarget 7, 14983-15017. Link
Kural, K.C., Tandon, N., Skoblov, M., Kel-Margoulis, O.V. and Baranova, A.V. (2016) Pathways of aging: comparative analysis of gene signatures in replicative senescence and stress induced premature senescence. BMC Genomics 17(Suppl 14), 1030. Link
Borlak, J., Singh, P. and Gazzana, G. (2015) Proteome mapping of epidermal growth factor induced hepatocellular carcinomas identifies novel cell metabolism targets and mitogen activated protein kinase signalling events. BMC Genomics 16, 124. Link
Shi, Y., Nikulenkov, F., Zawacka-Pankau, J., Li, H., Gabdoulline, R., Xu, J., Eriksson, S., Hedström, E., Issaeva, N., Kel, A., Arnér, E.S., Selivanova, G. (2014) ROS-dependent activation of JNK converts p53 into an efficient inhibitor of oncogenes leading to robust apoptosis. Cell Death Differ. 21, 612-623. Link
Schlereth, K., Heyl, C., Krampitz, A.M., Mernberger, M., Finkernagel, F., Scharfe, M., Jarek, M., Leich, E., Rosenwald, A., Stiewe, T. (2013) Characterization of the p53 Cistrome – DNA Binding Cooperativity Dissects p53’s Tumor Suppressor Functions. PLoS Genet. 9, e1003726.Link
Nikulenkov, F., Spinnler, C., Li, H., Tonelli, C., Shi, Y., Turunen, M., Kivioja, T., Ignatiev, I., Kel, A., Taipale, J., Selivanova, G. (2012) Insights into p53 transcriptional function via genome-wide chromatin occupancy and gene expression analysis. Cell Death Differ. 19, 1992-2002. Link
Zawacka-Pankau, J., Grinkevich, V.V., Hunten, S., Nikulenkov, F., Gluch, A., Li, H., Enge, M., Kel, A., Selivanova, G. (2011) Inhibition of glycolytic enzymes mediated by pharmacologically activated p53: targeting Warburg effect to fight cancer. J. Biol. Chem. 286, 41600-41615. Link
How to cite Match
Kel AE, Gössling E, Reuter I, Cheremushkin E, Kel-Margoulis OV, Wingender E. MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 2003;31(13):3576-3579. Link
