FOXM1 in multiple myeloma
TFClass 3.3.1.13.1
Transcription factor FOXM1 is known as one of the key regulators in multiple myeloma. A very recent publication in Oncogene has proven the mechanism of FOXM1 action, this is a positive regulation of myeloma cell metabolism, especially of glycolysis and oxidative phosphorylation (Cheng Y. et al., 2022, Oncogene, v.41: 3899-3911). The findings published therein allow to refer to FOXM1 as “a key driver of myeloma metabolism”.
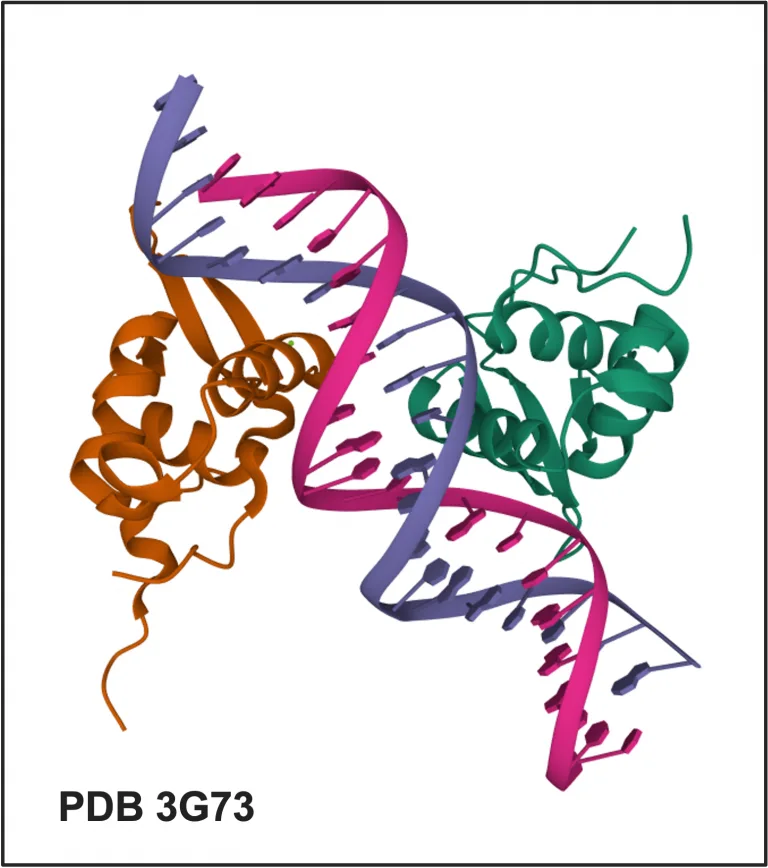
FOXM1 belongs to a big family of transcription factors, Forkhead box (FOX) factors. The FOX family of factors, together with E2F-related and RFX-related factor families, constitute a class of fork head / winged helix factors, which in turn belongs to the superclass of helix-turn-helix domains (Wingender at al., 2018; TFClass link).
FOXM1 is known for his role in more than 40 different tumors, among them are stomach, colorectal, ovarian, prostatic neoplasms; non-small-cell lung carcinoma, hepatocellular carcinoma, thyroid cancer and others. Disease similarity map for multiple myeloma can be viewed here.
Expression of FOXM1 protein in normal blood plasma cells, T-cells, B-cells, dendritic cells, granulocytes, monocytes is extremely low as compared with many other tissues and organs. The highest expression of FOXM1 protein is normally observed in thymus and testis (HumanPSD database, Human Protein Atlas v20).
As a transcription factor, FOXM1 binds to its specific binding sites in the promoters or enhancers of its target genes and regulates their transcription. In the TRANSFAC 2.0database, there are five positional weight matrices for FOXM1, either built from individual genomic binding sites or based on a ChIP-seq dataset. The core binding motif of FOXM1 is TRTTTR, where R can be G or A.
FOXM1 is known to regulate transcription of several functional groups of genes. Among them are cell cycle genes, CDC6 and CDC25A; genes encoding other transcription factors, ESR1, GATA3, SNAI1, SOX2, STAT3; growth factor genes VEGFA, PDGFA; glycolysis enzyme LDHA.
FOXM1 is well studied to be phosphorylated in a cell-cycle-dependent manner. In the TRANSPATH database, one can find details on FOXM1 phosphorylation by several cyclin dependent kinases, namely by the cdk:cyclin complexes CDK6:CycD3, CDK4:CycD1, CDK1:CycB as well as by polo-like kinase PLK1.
Many more details can be found in the integrated database TRANSFAC + TRANSPATH + HumanPSD, in the Locus Report for human FOXM1:
Open this report as a PDF file.
Cheng Y. et al., 2022. FOXM1 regulates glycolysis and energy production in multiple myeloma. Oncogene, 41, 3899-3911. PMID: 35794249
Wingender E. et al., 2018. TFClass: expanding the classification of human transcription factors to their mammalian orthologs. Nucleic Acids Res. 46, D343-D347. PMID: 29087517
This page was published and last revised on 16.08.2022
Examples of some other transcription factors in cancer can be found here.
