From Gene Lists to Biological Causes: Decoding the “Why” Behind Gene Expression Changes
Written by:

The Missing Question in Gene Expression Analysis:
Imagine sitting in a concert hall as a symphony orchestra plays.
Suddenly the music shifts — violins surge, brass intensifies, percussion accelerates. Dozens of instruments change at once.
You could carefully record which instruments became louder or quieter.
But that would not explain the music.
Because orchestras do not change by accident. Behind the sound stands the conductor, guiding the entire ensemble so that many instruments move together.
Gene expression works in much the same way.
When hundreds of genes change simultaneously, they are rarely acting alone. These coordinated changes are usually orchestrated by transcription factors and signaling pathways — the conductors of the cell’s regulatory symphony.
Understanding biology therefore requires more than listing changed genes.
It requires identifying the conductors behind the transcriptional music.
Over the past two decades, high-throughput technologies such as RNA-seq, microarrays, and multi-omics profiling have transformed biological research. Today, generating large datasets that describe changes in gene expression is relatively straightforward. A typical experiment may identify hundreds or even thousands of differentially expressed genes between two conditions—such as healthy versus diseased tissue, treated versus untreated cells, or young versus aged organisms.
However, this abundance of data introduces a fundamental challenge.
While differential expression analysis answers an important question — “What genes changed?” — it rarely explains the more important biological question: Why did these genes change?
Researchers frequently encounter the following problems:
- Long lists of upregulated and downregulated genes with no clear biological driver
- Multiple enriched pathways but no obvious causal mechanism
- Difficulty linking gene expression changes to upstream regulatory events
- Challenges identifying actionable targets or master regulators
In other words, gene expression data often provides symptoms of a biological process, but not the regulatory causes behind it.
Understanding these causes requires looking upstream in the regulatory hierarchy of the cell.
Gene expression changes are rarely random
In biological systems, coordinated changes in gene expression rarely occur by chance. Instead, they are usually controlled by regulatory networks composed of transcription factors, signaling pathways, and regulatory modules.
At the core of these networks are transcription factors (TFs) — proteins that bind specific DNA sequences and control the transcription of target genes.
A single transcription factor can regulate dozens or hundreds of genes simultaneously, creating coordinated expression programs. These programs govern fundamental biological processes including:
- cell differentiation
- immune responses
- stress signaling
- cancer progression
- aging and development
When a transcription factor becomes activated or inhibited, it can trigger widespread changes across the transcriptome.
Therefore, rather than focusing only on the genes that changed, researchers increasingly seek to identify:
- Which transcription factors caused these changes
- Which signaling pathways activated those regulators
- Which regulatory networks orchestrate the observed phenotype
This shift from gene lists to regulatory networks represents a major step toward mechanistic understanding of biological systems.
Moving upstream: From genes to regulatory networks
To identify the causes of gene expression changes, researchers must move upstream in the regulatory hierarchy.
This process is commonly known as upstream analysis.
Instead of starting with candidate regulators, upstream analysis works in the opposite direction:
- Begin with a list of differentially expressed genes.
- Identify transcription factor binding motifs enriched in their promoters.
- Predict transcription factors likely responsible for regulating those genes.
- Trace these regulators further upstream to signaling pathways and master regulators.
This approach transforms gene expression data from a descriptive dataset into a hypothesis-generating tool for regulatory biology.
A typical upstream analysis pipeline involves three major conceptual steps:
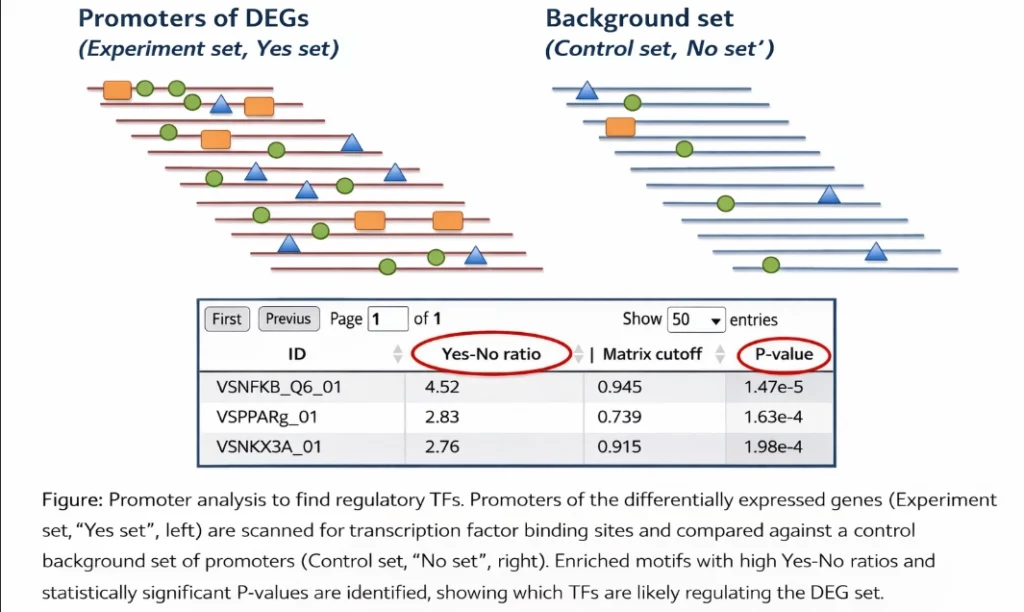
Step 1: Identifying regulatory signals in gene promoters
Genes are controlled through regulatory sequences in their promoters and enhancers where transcription factors bind. By analyzing these regulatory regions, researchers can detect overrepresented transcription factor binding motifs among differentially expressed genes.
If a particular motif appears frequently in the promoters of upregulated genes, it suggests that the corresponding transcription factor may be driving the observed expression pattern.
This step converts a gene list into a set of candidate transcriptional regulators.
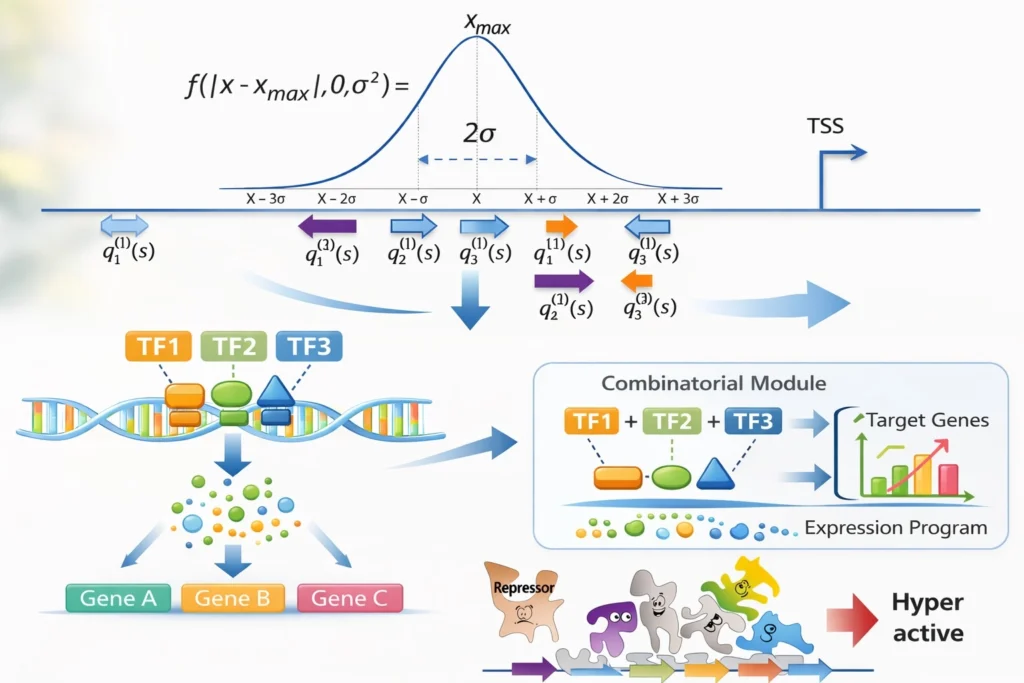
Step 2: Identifying transcription factor combinations
In reality, gene regulation rarely depends on a single transcription factor.Instead, transcription factors often act in combinatorial modules, where multiple regulators cooperate to control gene expression. Identifying these regulatory modules provides deeper insight into the regulatory architecture behind the data.
Such modules can reveal:
- cooperative transcription factor activity
- condition-specific regulatory programs
- context-dependent gene regulation
Understanding these regulatory combinations helps explain how specific gene expression patterns emerge.
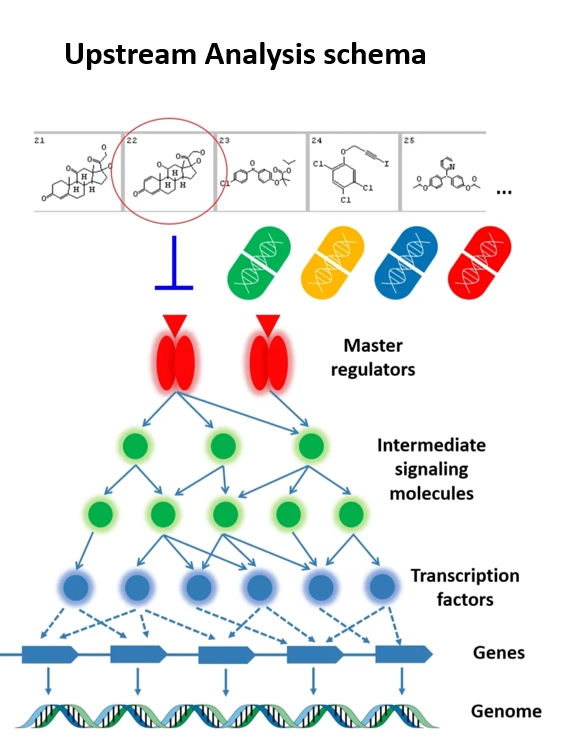
Step 3: Tracing signaling pathways to master regulators
Once transcription factors are identified, the next step is to determine what activated them.
Transcription factors are typically controlled by signaling pathways that respond to external or internal cellular stimuli.
By tracing signaling networks upstream of the predicted transcription factors, researchers can identify master regulators — key molecules whose activity ultimately drives the transcriptional response.
Master regulators often represent:
- signaling hubs
- disease drivers
- potential therapeutic targets
This final step provides the missing link between gene expression changes and cellular signaling mechanisms.
Case study: Upstream analysis in triple-negative breast cancer
To illustrate how upstream analysis works in practice, consider an example from cancer biology. Triple-negative breast cancer (TNBC) is an aggressive breast cancer subtype characterized by the absence of estrogen receptor, progesterone receptor, and HER2. Because these common therapeutic targets are missing, identifying the molecular drivers of TNBC gene expression and growth remains a major research focus.
In this example, an RNA-seq dataset (NCBI GEO: GSE188914) was analyzed, comparing a TNBC cell line (MDA-MB-231) with an ER-positive breast cancer cell line (MCF-7). The objective was to identify the master regulators responsible for the distinct transcriptional behavior of TNBC cells.
The dataset contains gene expression profiles from three replicates of each cell line. Using the geneXplain platform, integrating curated knowledge from the TRANSFAC® and TRANSPATH® databases, the analysis aimed to uncover the upstream regulatory mechanisms driving TNBC-specific gene expression patterns.
The analysis proceeded through several stages.
Differential gene expression analysis
The study first identified genes whose expression levels differed significantly between TNBC samples and control tissues.
This produced a list of genes associated with processes such as:
- cell proliferation
- immune signaling
- extracellular matrix remodeling
- inflammatory pathways
However, as with many transcriptomic studies, the gene list alone did not reveal the regulatory causes of these changes.
Promoter and transcription factor analysis
Researchers then analyzed the promoter regions of these genes to detect enriched transcription factor binding sites.
This analysis revealed several transcription factors likely responsible for regulating the observed expression changes.
These included regulators associated with:
- inflammatory signaling
- stress response pathways
- tumor progression
Such findings suggested that the TNBC transcriptional program may be controlled by specific transcription factor networks rather than isolated gene changes.
Identification of master regulators
The final step traced signaling pathways upstream of the identified transcription factors.
This revealed several master regulatory molecules capable of activating the regulatory network observed in the TNBC dataset.These master regulators represent potential control points in the disease network, offering valuable insights into mechanisms that may drive tumor progression.
Results and Discussion
To understand the regulatory mechanisms distinguishing triple-negative breast cancer (TNBC) from ER-positive breast cancer cells, promoter and upstream pathway analyses were performed on the set of genes upregulated in TNBC.
Promoter Analysis: Identifying Candidate Transcription Factors
Promoter analysis identified 68 significantly enriched DNA motifs in the promoters of TNBC-upregulated genes. These motifs correspond to 132 transcription factors (TFs), reflecting the fact that multiple TFs can share similar DNA binding patterns.
Interestingly, 46 of these TFs were themselves upregulated in the TNBC expression dataset (from the list of 437 genes), suggesting the presence of a self-reinforcing regulatory network in which activated TFs regulate additional genes and potentially sustain each other’s expression.
One notable example is IRF1 (Interferon Regulatory Factor 1). The IRF1 gene was strongly upregulated in TNBC cells (log₂ fold change ≈ +3.0), and its binding motif was ~2.2-fold enriched in promoters of TNBC-upregulated genes. The analysis predicted 54 potential IRF1 target genes within this gene set, indicating that IRF1 may play a central role in driving TNBC-specific transcriptional programs.
Other enriched motifs corresponded to regulators such as the E2F family, AP-1 complex, NF-κB, and the glucocorticoid receptor (GR)—all transcription factors known to regulate cell proliferation, inflammation, stress responses, and cancer progression. Together, these findings provide a preliminary list of candidate transcriptional regulators responsible for the altered gene expression observed in TNBC cells.
Pathway Analysis: Discovering Master Regulators
To identify upstream drivers of these transcription factors, the candidate TFs were subjected to upstream network analysis using signaling pathway data. This step identified 12 master regulator candidates located at convergence points of signaling pathways controlling the TF network.
Several of these regulators were also upregulated in TNBC cells, strengthening their potential functional relevance.
Key master regulators included:
- CD40, a TNF-superfamily receptor, emerged as the top candidate and was nearly 10-fold upregulated in TNBC cells. CD40 signaling can activate NF-κB and inflammatory pathways, which may contribute to the inflammatory gene expression signature often observed in TNBC.
- Glucocorticoid receptor (GR / NR3C1) was ~4.5-fold upregulated, suggesting enhanced glucocorticoid signaling that may influence immune responses, metastasis, or therapy resistance.
- E2F1 and E2F7, both significantly upregulated (~2.9-fold and ~3.6-fold respectively), highlight strong cell-cycle deregulation in TNBC.
- NEK2A, a mitotic kinase involved in centrosome regulation, was ~2.4-fold upregulated, indicating active mitotic signaling networks.
- c-Myc, a global oncogenic transcriptional regulator (~1.7-fold upregulated), likely contributes to the widespread transcriptional amplification seen in aggressive cancer cells.
- Components of the NF-κB pathway were also identified, supporting the presence of constitutive inflammatory signaling in TNBC.
Biological Interpretation and Implications
Taken together, the analysis suggests that TNBC cells are driven by a coordinated regulatory network combining inflammatory signaling (CD40–NF-κB), stress hormone signaling (GR), and deregulated cell-cycle control (E2F, Myc, NEK2). These master regulators likely activate downstream transcription factors and gene expression programs that shape the aggressive phenotype of TNBC.
Importantly, many of these regulators represent known drug targets or biomarkers, providing valuable hypotheses for further experimental validation. For example, inhibiting CD40 signaling, glucocorticoid receptor activity, Myc function, or mitotic kinases could help determine their role in sustaining the TNBC transcriptional program.
While upstream analysis does not prove causation, it significantly narrows the search for causal regulators, guiding researchers toward targeted experiments such as knockdown or pharmacological inhibition studies to validate these predicted drivers of TNBC biology.
Conclusion: From data to biological insight
As genomic datasets continue to grow in size and complexity, the challenge facing researchers is no longer data generation, but data interpretation.
Understanding biological systems requires connecting observed molecular changes to the regulatory networks that control them.
By shifting focus from “what changed” to “why it changed,” researchers can move closer to uncovering the true mechanisms underlying cellular behavior, disease progression, and therapeutic response.
Regulatory network analysis provides a framework for making this transition — transforming gene expression datasets into meaningful biological insight.
References
- Kel, A. et al. TRANSFAC and its module-based analysis tools. Nucleic Acids Research.
- Wingender, E. et al. TRANSFAC: transcription factor binding site database. Nucleic Acids Research.
- Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. PNAS.
- Hanahan, D., Weinberg, R. Hallmarks of cancer: the next generation. Cell.
- Upstream analysis methodology and case study in triple-negative breast cancer:
https://genexplain.com/upstream-analysis-identifying-master-switches-in-gene-regulation/
