About TRANSFAC
All three packages — Basic, Pathways, and Diseases — include the TRANSFAC database, giving you direct access to expertly curated regulatory knowledge from day one.
TRANSFAC BASIC
Discover TF binding sites in promoters and enhancers of your genes
TRANSFAC® is the database of eukaryotic transcription factors, their genomic binding sites and DNA-binding profiles. Dating back to a very early compilation 35 years ago, it has been carefully maintained and curated to become the gold standard in the field. Since then this biggest collection of transcription factors and their genomics binding sites has served the scientific community as the most reliable and comprehensive resource for gene regulation studies.
You can use TRANSFAC® as an encyclopedia of transcriptional regulation, or as a tool to identify potential TFBSs by applying its library of positional weight matrices, a unique collection of DNA-binding models. The latter can be done with the included MATCH (Suite) tools or with any of the respective modules in the geneXplain platform.
TRANSFAC database
It’s been almost 35 years since the date of the first TRANSFAC® database publication, and since then this biggest collection of transcription factors and their genomics binding sites has served the scientific community as the most reliable and comprehensive resource for gene regulation studies.
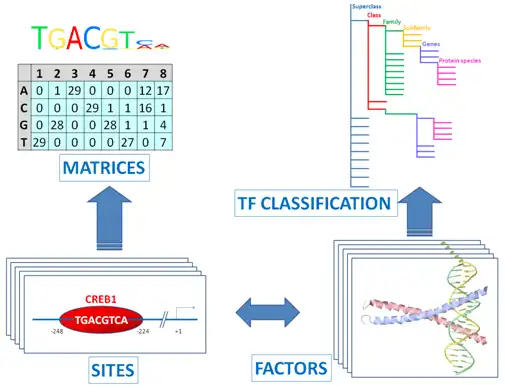
The core of TRANSFAC® comprises contents of two domains: One documents TF binding sites, usually in promoters or enhancers. The other describes the binding proteins (TFs).
Transcription factors are subsumed to classes, based on the general properties of their DNA-binding domains. This early attempt has been expanded to a comprehensive TF classification, the latest version of which can be found here.
Here is a general introduction to the online TRANSFAC® database. It shows how to operate in the basic interface and perform searches in the TRANSFAC® database.)
TRANSFAC® includes information on over 49,000 transcription factors and their genomics binding site models (PWMs – positional weight matrices) for a variety of eukaryotic species.
Positional Weight Matrices
Binding sites referring to the same TF are merged into a positional weight matrix. Such a matrix reflects the frequency with which each nucleotide is found in each position of this TF’s binding sites and, thus, the base preference in each position.
Get free reports and case studies to your Email
Here is an example of PWM for binding sites for E2F-1 transcription factors.
TRANSFAC® includes over 10,000 Positional Weight Matrices (PWMs) that are robust models for predicting TF binding sites in genomes of various animals, plants, and fungi.
Experimentally proven TF binding sites
TRANSFAC® includes over 50,000 experimentally proven TF binding sites and about 100 million ChIP-seq TF binding regions.
Example reports
Here is an example of a report on binding sites for CREB1 TF in a promoter of human CD14 gene.
Promoters, Enhancers
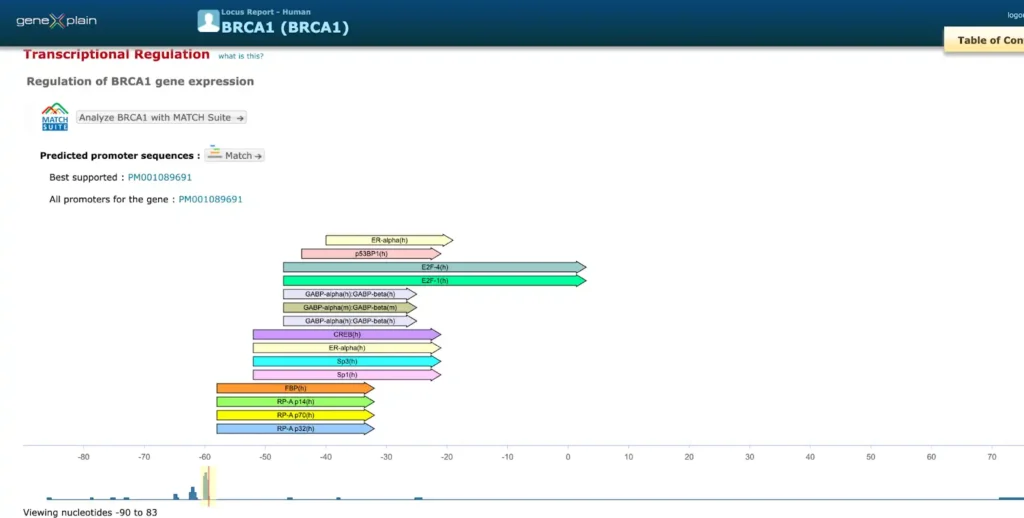
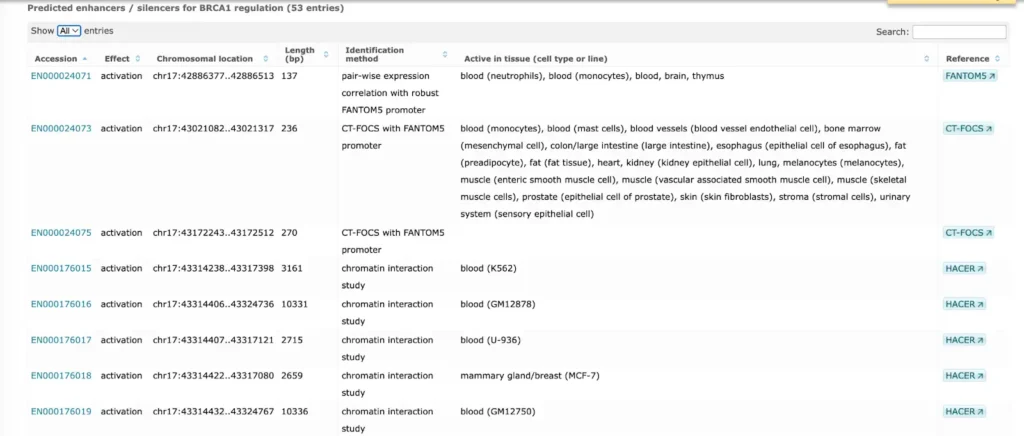
TRANSFAC® includes promoter genomic annotation for various vertebrate and invertebrate species as well as for plants and fungi. TRANSFAC® also includes genomic information for over 200,000 human enhancers and silences acting in different tissues and cell type-specific.
Example:
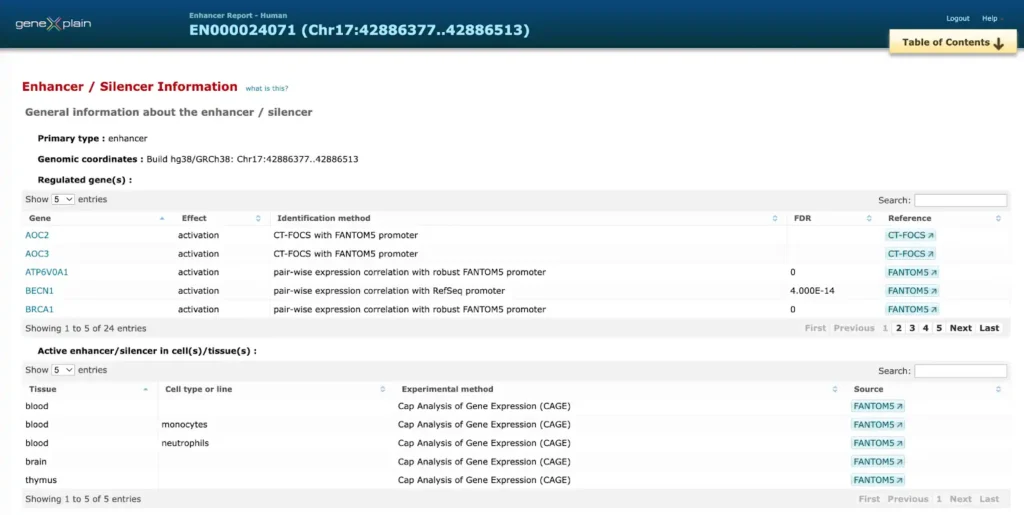
Here is an example of a report on promoter and enhancer annotation of the human BRCA1 gene
The enhancer and promoter data is accumulated from numerous public repositories of CAGE, FANTOM and Hi-C data in bulk and single cell.
Site search and site enrichment tools
TRANSFAC® includes one of the best site search and site enrichment tools equipped with powerful user interfaces for application in different analysis tasks.
TFBS prediction in the TRANSFAC database interface (MATCH, FMatch, CMsearch)
The TRANSFAC® database provides a wide variety of tools for predicting transcription factor binding sites (TFBS) in the studied DNA sequences. In the portal interface of the TRANSFAC® database you will find the Match, FMatch, and Composite Model analysis tools for TFBS prediction under the Tools menu button:
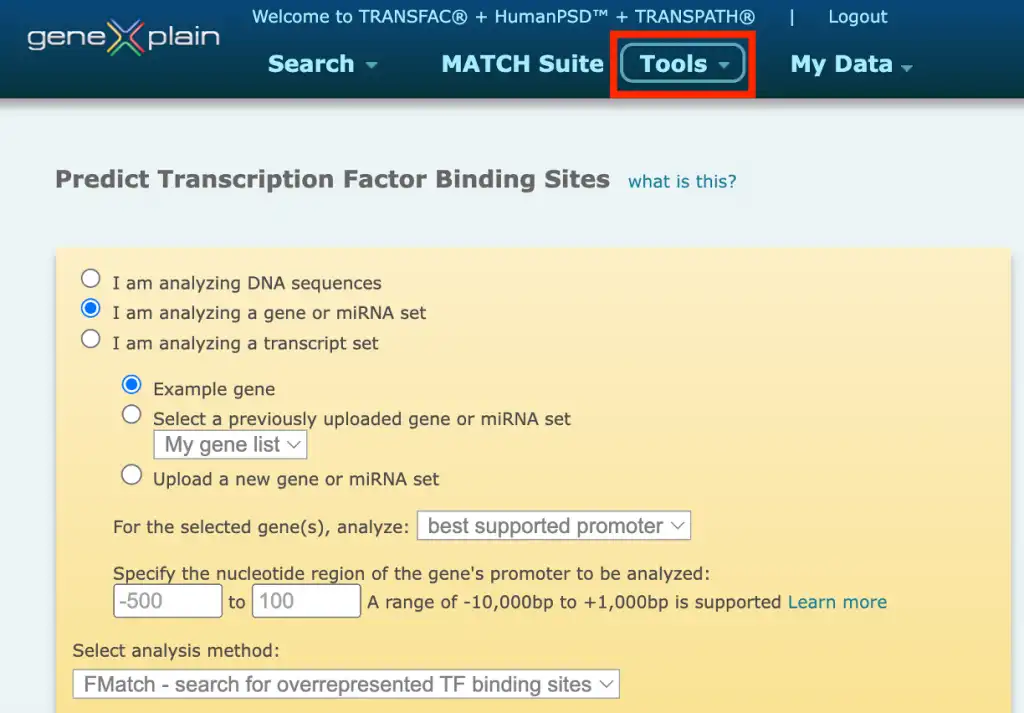
1. Match – search for TF binding sites
This option uses the Match algorithm, in combination with a selected profile containing a list of matrices and their assigned cut-offs to search for individual transcription factor binding sites that meet the specified cut-offs.The Match option is recommended when the broadest set of results is desired.
2. Composite model – search by pairs of TFs
This option uses the Composite Model algorithm, in combination with a selected model or models which represent pairs of transcription factors known to act together to coordinately control gene regulation, and their assigned cut-offs to search for pairs of transcription factor binding sites that meet the specified cut-offs.The Composite Model option is recommended when specific information about coordinate regulation is known, or when more stringent results are desired.
3. FMatch – search for overrepresented TF binding sites
This option is used to find sites which are overrepresented in a set of analyzed sequences (e.g. promoters from differentially expressed genes or ChIP-Seq fragments) in comparison to a background set (e.g. promoters from genes whose expression did not change under the same conditions or random sequences).
Predicting transcription factor binding sites in a DNA sequence involves several steps:
Input Formats
Match in the geneXplain portal interface accepts the following DNA sequence formats as input:
FAST
GenBank
EMBL
RAW
Alternatively, a new sequence via genomic coordinates in the .bed format can be uploaded. This input format is supported for human (hg38/GRCh38), mouse (mm39/GRCm39), rat (rn6/RGSC 6.0), pig (Sscofa11.1), macaque (Mmul8.0.1), Arabidopsis (TAIR10), and fruit fly (BDGP6) genomes.
The geneXplain portal Match tool also accepts gene or miRNA set as the input for TFBS prediction analysis.
Еhe option to upload a list of genes or miRNAs for binding site prediction is available for those organisms, for which promoter sequences are provided in TRANSFAC database, namely: Human, Mouse, Rat, Arabidopsis thaliana, Soybean, Rice, Pig, Macaca mulatta, Drosophila, Dog, Chimpanzee, and Plasmodium falciparum 3D7.
Example Reports
The results of Match, FMatch and Composite Model analysis are presented in the respective analysis report which is comprised of three sections:
(Click on the section for detailed information)
Analysis summary
The Analysis summary section provides an overview of the count of sequences analyzed, the number of sites found, etc. In the FMatch Analysis Report the Analysis Summary contains a summary on the Experimental data set, as well as a summary on the Background data set.
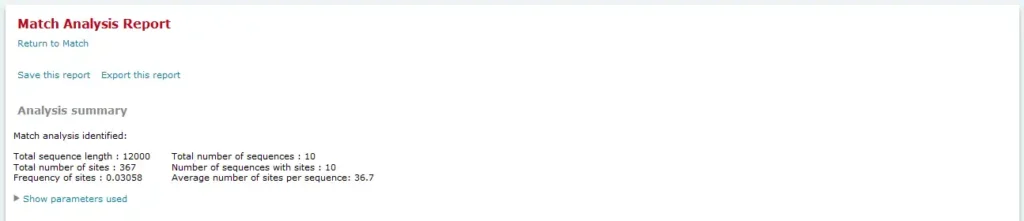

Matrix summary (Match, FMatch), Model summary (Composite Model)
For Match results, the Matrix summary section provides an overview of the matrices for which at least one binding site was predicted:
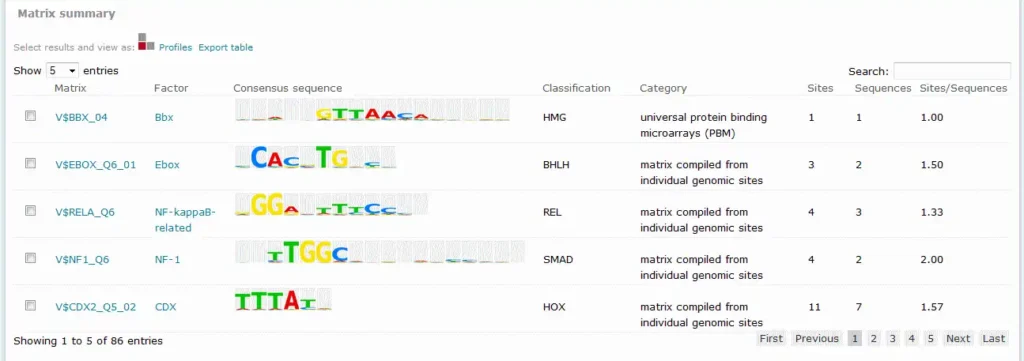
When available, additional info is provided on the experimental support of the possible biological connection between the factor and the gene. The total number of binding sites predicted for the matrix across all sequences within the analysis is shown in the Sitescolumn on the Matrix summary table.
The Sequences column refers to the number of sequences within the sequence set for which at least one binding site was predicted for the matrix. The value in the Sites per sequence column provides the average number of binding sites predicted for the matrix per sequences within the sequence set.
For Composite Model results, the Model summary section provides an overview of results for each model considered in the analysis:
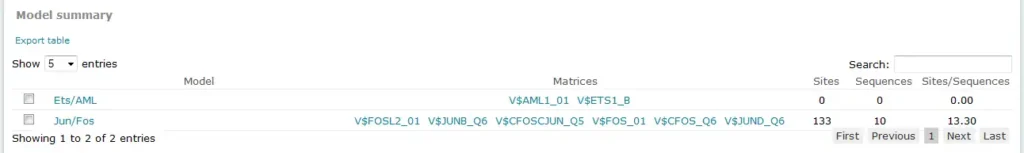
The Sites column in this table provides the count of total binding sites predicted for the model across all sequences within the analysis. The Sequences column provides the count of sequences within the sequence set for which at least one binding site was predicted for the model. The Sites per sequence column provides the average number of binding sites predicted for the model per sequences within the sequence set.
For FMatch results, the Matrix summary section provides an overview of the matrices for which at the optimized cut-offs the over- or underrepresentation of sites in the experimental data set versus the background data set fit the p-value threshold:
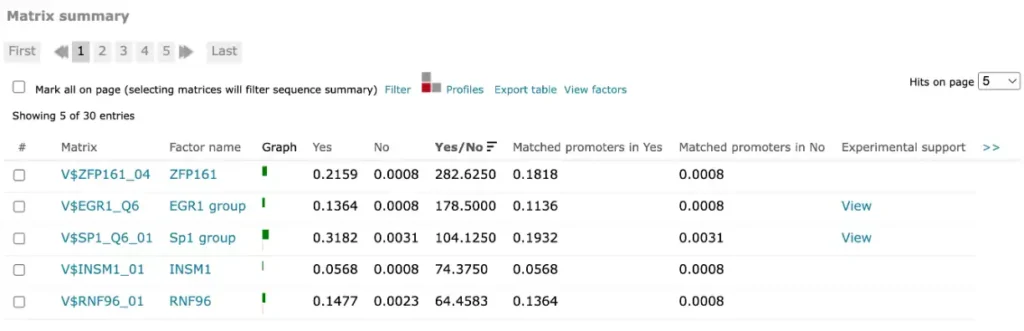
The Graph column displays the relative number of sites for the selected matrix in the experimental data set (green bar) versus in the background set (red bar). The Yes and No columns contain the relative number of sites for the selected matrix in the experimental data set and the relative number of sites for the selected matrix in the background data set respectively.
The Yes/No ration column shows the relative number of sites for the selected matrix in the experimental data set divided by the relative number of sites in the background data set. The Matched promoters in Yes and Matched promoters in No columns show the relative number of sequences/promoters in the experimental data set with at least one site for the selected matrix and the relative number of sequences/promoters in the background data set with at least one site for the selected matrix respectively.
Sequence summary
For Match and FMatch results, the Sequence summary section provides, for each sequence analyzed, a graphical display of the predicted binding sites and a tabular summary. In the Sequence Summary of the FMatch Report, you can switch between the Experimental set and the Background set.
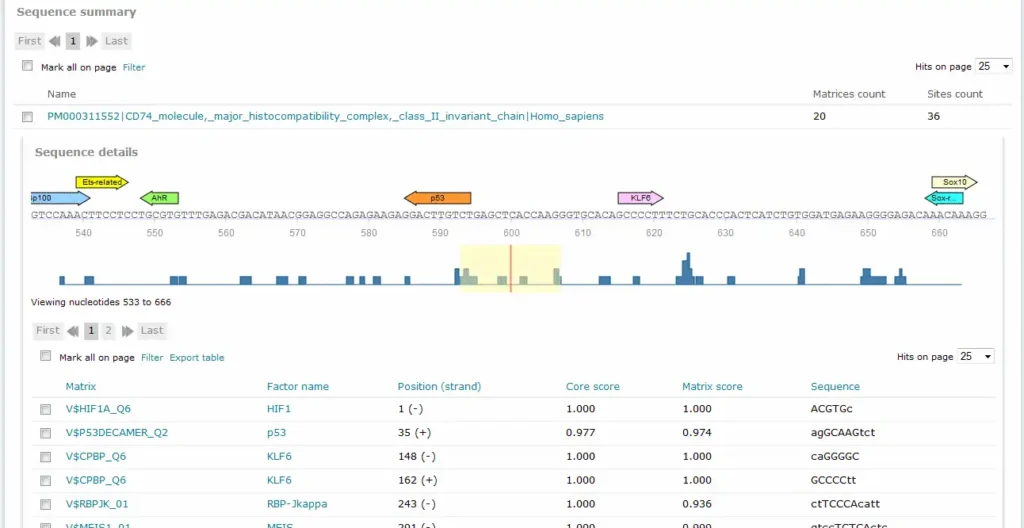
If more than one sequence was submitted for analysis, a click on the sequence name would view the binding sites graphical display and tabular summary. If only one sequence was submitted for analysis, the view will open automatically.
The Position (strand) column indicates the starting position of the match in the input sequence and the strand, (+) or (-), on which it can be found. In case of analyzed genomic intervals or promoter sequences submitted from TRANSFAC, genomic coordinates are provided. The Core score column indicates the score for core similarity (core match). The Matrix score column indicates the score for matrix similarity (matrix match). The Sequence column identifies the portion of the input sequence that was identified as the binding site. Capital letters indicate the positions in the sequence that match with the core sequence of the matrix, while the lower case letters refer to positions which match to other parts of the matrix. When available, experimental support info is provided, which lists supporting lines of evidence from the scientific literature supporting a possible biological connection between the factor and the gene.
In the graphical output, the predicted sites are shown as arrows above the respective part of the sequence with the factor name printed within the arrow:

A click on the arrow will open a pop-up window with the binding site information summary and a hyperlink to the corresponding Matrix Report.
When the genomic coordinates of the analyzed sequences are known either via .bed coordinate upload or via the use of stored TRANSFAC promoters, the binding sites predicted with Match, CMsearch or FMatch can be filtered by intervals from the database of ChIP fragments (Binding fragments for transcription factors from ChIP-seq or similar experiments), DNase hypersensitivity sites or Phastcons intervals (intervals of conservation, as determined by 46-way phastcons and 60-way phastcons placental mammals tracks for human and mouse at UCSC ). The location of the selected intervals/features is displayed by color-coded bars underneath the analyzed sequence and gray lines in the frequency bar. When one of these features is selected, all hits outside the respective intervals are excluded from the result. When more than one feature is selected at the same time, all hits outside the intersection of the intervals are excluded from the result. For the filtering, the matrix (and model) hits are allowed to extend up to three nucleotides outside the intervals.
For Composite Model results, the Sequence summary section provides, for each sequence analyzed, a graphical display of the predicted binding sites and a tabular summary:
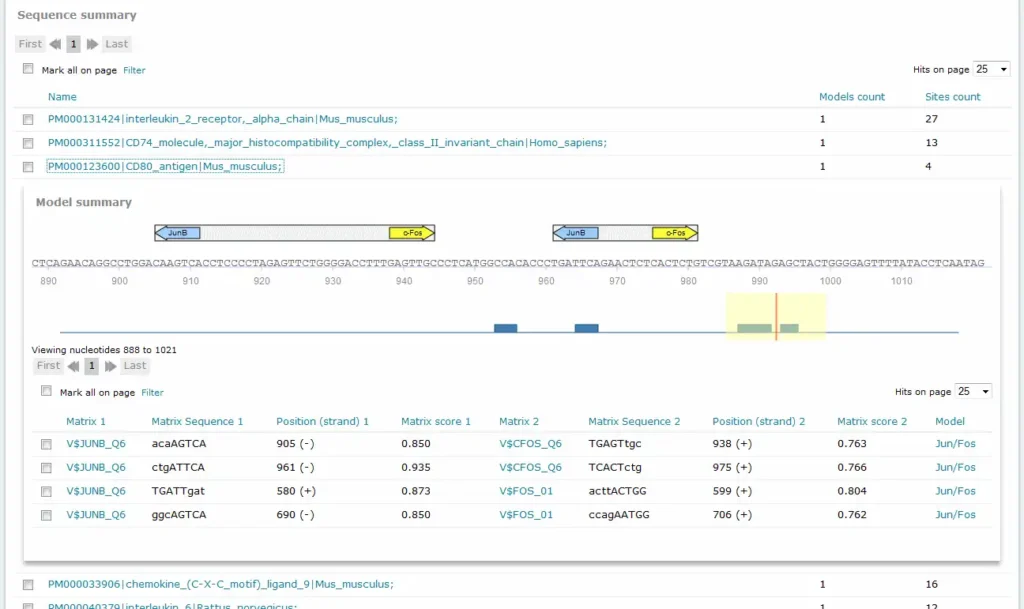
If more than one sequence was submitted for analysis, a click on the sequence name would view the binding sites graphical display and tabular summary. If one sequence was submitted for analysis, the view will open automatically.
The columns Matrix 1 and Matrix 2 identify the respective matrix and either provide a hyperlink to the corresponding TRANSFAC Matrix Report or, in the case of user-defined matrix, display the name of the user matrix. Columns Sequence 1 and Sequence 2 identify the matching sequence. Capital letters indicate the positions in the sequence that match with the core sequence of the matrix, while the lower case letters refer to positions which match to other parts of the matrix. Columns Position (strand 1) and Position (strand) 2 indicate the starting position of the match in the input sequence and the strand, (+) or (-), on which it can be found. The columns Matrix score 1 and Matrix score 2 indicate the score for matrix similarity (matrix match). The column Model provides the name of the model.
The obtained results can be saved to the geneXplain portal cloud storage using the “Save this report” link at the top of the analysis report, or dowloaded to the local computer using the “Export this report”.
More information on TFBS search in the geneXplain portal interface can be found in the “Gene Regulation Analysis Tools” –> “Predicting TF-Binding Sites” section of the portal user guide.
TRANSFAC Features & Benefits
TRANSFAC: Your gateway to precision gene regulation
TRANSFAC isn’t just another database—it’s the original, the one everyone else imitates.
When precision, clarity, and actionable insights from genomic data matter, TRANSFAC is your best ally.
Motifs & TF-Binding Sites: Precision Defined
10,000+ Curated PWMs
Access the largest, highest-quality library of eukaryotic transcription factor motifs
Instant Variant Impact
Quickly see how genomic variants impact transcription factor binding, streamlining mutation interpretation

AI-Enhanced Predictions
Accurately predict TF interactions, saving time and increasing confidence in your results
Key outcomes:
Eliminate guesswork in genomic interpretation, dramatically accelerating research timelines
Unique features:
Unlike simplified motif databases, TRANSFAC’s PWMs are extensively validated, offering far superior reliability
Promoters & Enhancers: Explore Deeper, Find More
Experimentally Validated Data
Leverage the largest collection of experimentally proven TF-gene interactions for robust insights

Discover Hidden Regulators
Detect complex regulatory interactions others overlook, including synergistic composite elements
Rapid Target Identification
Effortlessly pinpoint precise targets tailored to specific tissues or conditions
Key outcomes:
Quickly transition from raw data to validated hypotheses, improving research efficiency and accuracy
Unique features:
While others rely on computational predictions alone, TRANSFAC provides unmatched experimental validation
Pathways & Master Regulators: Connect the Biological Dots
Visual Network Clarity
Simplify complex data using over 1.2 million expert-validated molecular interactions
Custom Network Creation
Build personalized pathways effortlessly, tailored specifically to your research questions
Uncover Master Regulators
Identify critical upstream drivers missed by traditional approaches, highlighting impactful therapeutic targets
Key outcomes:
Move beyond surface-level pathway analysis to deeply understand disease mechanisms
Unique features:
TRANSFAC integrates uniquely curated pathway data, far exceeding standard pathway tools
Multi-Omics: Integration Without Complexity
Automated Analysis Pipeline
Easily integrate genomics, transcriptomics, proteomics, epigenomics, and metabolomics data
Advanced Upstream Analysis
Precisely identify master transcriptional regulators from complex multi-omics datasets

Key outcomes:
Streamline multi-omics analyses, drastically reducing time from data to actionable insights
Unique features:
Unlike fragmented analysis tools, TRANSFAC provides comprehensive, fully automated multi-omics integration
Biomarkers & Drug Discovery: Faster Insights, Better Outcomes
Extensive Biomarker Database
Over 140,000 curated gene-disease and drug-target associations
Disease Mechanism Reconstruction
Effortlessly reconstruct molecular drivers behind diseases, enhancing therapeutic strategies
Key outcomes:
Rapidly discover biomarkers and drug targets, greatly improving drug development timelines
Unique features:
TRANSFAC’s biomarker data depth and disease mechanism insights surpass competitors significantly
Precision Medicine: Truly Personalized, Highly Effective
Personalized Therapeutic Insights
Quickly identify patient-specific drug targets from individual multi-omics profiles
Drug Repurposing Excellence
Efficiently explore off-label and novel drug combinations for personalized care
Key outcomes:
Achieve truly personalized treatment strategies, greatly improving patient outcomes
Unique features:
TRANSFAC uniquely integrates omics data to deliver actionable personalized therapeutic insights
Powerful, User-Friendly Workflows
200+ Ready-to-Use Pipelines
Effortlessly execute complex analyses without coding
Visual Pipeline Customization
Intuitively tailor your analyses with visual programming
Flexible APIs & Visualization Tools
Seamlessly manage workflows and visualize results intuitively
Key outcomes:
Remove bioinformatics bottlenecks, empowering biologists to directly drive data analysis
Unique features:
TRANSFAC’s user-friendly workflows eliminate the steep learning curves typical of competitor platforms
Why Choose TRANSFAC? Simply Unmatched.
35 Years of Expertise
Meticulous curation ensures unparalleled accuracy and trustworthiness

Largest Validated Dataset
The gold standard for experimentally proven TF binding sites
Robust AI Integration
Advanced algorithms deliver exceptionally precise regulatory predictions
Comprehensive Systems Biology Approach
Seamlessly integrates transcription factors, pathways, and disease mechanisms
TRANSFAC:
Precision, Power, and Proven Insights for Gene Regulation.
TRANSFAC Applications in Research — Selected Publications
Novikova S., Tolstova T., Kurbatov L., Farafonova T., Tikhonova O., Soloveva N., Rusanov A., Zgoda V. (2024) Systems Biology for Drug Target Discovery in Acute Myeloid Leukemia. Int. J. Mol. Sci. 25(9), 4618 Link
Ivanov, S. M., Tarasova, O. A., & Poroikov, V. V. (2023). Transcriptome-based analysis of human peripheral blood reveals regulators of immune response in different viral infections. Frontiers in immunology, 14, 1199482. Link
Menck, K., Heinrichs, S., Wlochowitz, D., Sitte, M., Noeding, H., Janshoff, A., Treiber, H., Ruhwedel, T., Schatlo, B., von der Brelie, C., Wiemann, S., Pukrop, T., Beißbarth, T., Binder, C., & Bleckmann, A. (2021). WNT11/ROR2 signaling is associated with tumor invasion and poor survival in breast cancer. Journal of experimental & clinical cancer research : CR, 40(1), 395. Link
Andreev-Andrievskiy, A. A., Zinovkin, R. A., Mashkin, M. A., Frolova, O. Y., Kazaishvili, Y. G., Scherbakova, V. S., Rudoy, B. A., & Nesterenko, V. G. (2021). Gene Expression Pattern of Peyer’s Patch Lymphocytes Exposed to Kagocel Suggests Pattern-Recognition Receptors Mediate Its Action. Frontiers in pharmacology, 12, 679511. Link
Ivanov, S., Filimonov, D., & Tarasova, O. (2021) A computational analysis of transcriptional profiles from CD8(+) T lymphocytes reveals potential mechanisms of HIV/AIDS control and progression. Comput Struct Biotechnol J. 19, 2447–2459. Link
Li, X., Shi, J., & Li, L. M. (2025) The human intelligence evolved from proximal cis-regulatory saltations. Quantitative Biology 13(2) Link
Raj Murthi, S., Petry, A., Shashikadze, B., Stöckl, J. B., Schmid, M., Santamaria, G., Klingel, K., Kračun, D., Chen, X., Bauer, S., Schmitt, J. P., Flenkenthaler, F., Gorham, J., Toepfer, C. N., Potěšil, D., Hruška, P., Zdráhal, Z., Mayer, Z., Klop, M., Lehmann, L., … Wolf, C. M. (2025). Contribution of hypoxia-inducible factor 1alpha to pathogenesis of sarcomeric hypertrophic cardiomyopathy. Scientific reports, 15(1), 2132. Link
Cihan, M., Schmauck, G., Sprang, M., & Andrade-Navarro, M. A. (2025). Unveiling cell-type-specific microRNA networks through alternative polyadenylation in glioblastoma. BMC biology, 23(1), 15. Link
Jian, Y., Xu, H., Wang, Z., Zhang, Z., & Zhang, X. (2025). Histone modification-based functional characterization and genetic association of polymorphisms in LRRC6 and MTMR10 within CRC susceptibility regions 8q24 and 15q13.3. Gene, 943, 149286. Link
Ferreira Dos Santos, T. C., Silva, E. N., Frezarim, G. B., Salatta, B. M., Baldi, F., Simielli Fonseca, L. F., De Albuquerque, L. G., Magalhães Muniz, M. M., & Dos Santos Silva, D. B. (2025). Cis-eQTL analysis reveals genes involved in biological processes of the immune system in Nelore cattle. Gene, 937, 149138. Link
Hosseinpouri, A., Sadegh, K., Zarei-Behjani, Z., Dehghan, Z., & Karbalaei, R. (2025). Identification of critical genes and drug repurposing targets in entorhinal cortex of Alzheimer’s disease. Neurogenetics, 26(1), 27. Link
Woelk, J., Hornsteiner, F., Aschauer-Wallner, S., Stoitzner, P., Baier, G., & Hermann-Kleiter, N. (2025). Regulation of NK cell development, maturation, and antitumor responses by the nuclear receptor NR2F6. Cell death & disease, 16(1), 77. Link
Batool, F., Shireen, H., Malik, M. F., Abrar, M., & Abbasi, A. A. (2025). The combinatorial binding syntax of transcription factors in forebrain-specific enhancers. Biology open, 14(2), BIO061751. Link
Liang, Y., Yao, X., Han, J., Wang, J., Zhang, X., Zhao, D., Jiang, C., Geng, L., Lv, S., Liu, Z., & Mu, Y. (2024). Establishment of a CRISPR-Based Lentiviral Activation Library for Transcription Factor Screening in Porcine Cells. Animals : an open access journal from MDPI, 15(1), 19. Link
Dong, J. P., Xu, Y. C., Jiang, Y. N., Jiang, R. Z., Ma, L., Li, X. Z., Zeng, W. H., & Lin, Y. (2024). Identification of transcriptional signature change and critical transcription factors involved during the differentiation of mouse trophoblast stem cell into maternal blood vessel associated trophoblast giant cell. Cellular signalling, 123, 111359. Link
Mandić, K., Milutin Gašperov, N., Božinović, K., Dediol, E., Krasić, J., Sinčić, N., Grce, M., Sabol, I., & Barešić, A. (2024). Integrative analysis in head and neck cancer reveals distinct role of miRNome and methylome as tumour epigenetic drivers. Scientific reports, 14(1), 9062. Link
Syed, R., Rengasamy, P., Rajagopalan, S., Deiuliis, J. A., & Maiseyeu, A. (2024). MicroRNA 223 Enhances ABCA1 Protein Stability and Supports Efflux in Cholesterol-Burdened Macrophages. Cell Link
Nadeem, H., Jamal, S. B., Basheer, A., Bakhtiar, S. M., Faheem, M., Aziz, T., Nabi, G., Al-Harbi, M., & Raza, R. Z. (2024). Genetic Insights into Facial Variation and Craniofacial Development: Unraveling the Interplay of Genes, Expression Patterns, and Evolutionary Significance. Molecular biotechnology, 10.1007/s12033-024-01349-6. Advance online publication. Link
Zhou, Z., Lv, Y., Li, L., Yuan, X., Zhou, X., & Li, J. (2024). FoxO1 Mediated by H3K27me3 Inhibits Porcine Follicular Development by Regulating the Transcription of CYP1A1. Animals : an open access journal from MDPI, 14(23), 3514. Link
Zhou, L., Yang, Y., Qiao, Q., Mi, Y., Gan, Y., Zheng, Y., Wang, Y., Liu, M., & Zhou, Y. (2024). AKT1-Mediated NOTCH1 phosphorylation promotes gastric cancer progression via targeted regulation of IRS-1 transcription. Journal of cancer research and clinical oncology, 151(1), 15. Link
Zhou L., Yang Y., Ye Y., Qiao Q., Mi Y., Liu H., Zheng Y., Wang Y., Liu M., Zhou Y. (2024). Notch1 signaling pathway promotes growth and metastasis of gastric cancer via modulating CDH5. Aging (Albany NY). 16(16):11893–11903. Link
Li F., Wang J., Li M., Zhang X., Tang Y., Song X., Zhang Y., Pei L., Liu J., Zhang C., Li X., Xu Y., Zhang Y. (2024). Identifying cell type-specific transcription factor-mediated activity immune modules reveal implications for immunotherapy and molecular classification of pan-cancer. Brief Bioinform. 25(5):bbae368. Link
Lleshi E., Milne-Clark T., Yu H.L., Martin H.W., Hanson R., Lach R., Rossi S.H., Riediger A.L., Görtz M., Sültmann H., Flewitt A., Lynch A.G., Gnanapragasam V.J., Massie C.E., Dev H.S. (2024). Prostate cancer detection through unbiased capture of methylated cell-free DNA. iScience. 27(7):110330. Link
Ceroni F., Cicekdal M.B., Holt R., Sorokina E., Chassaing N., Clokie S., Naert T., Talbot L.V., Muheisen S., Bax D.A., Kesim Y., Kivuva E.C., Vincent-Delorme C., Lienkamp S.S., Plaisancié J., De Baere E., Calvas P., Vleminckx K., Semina E.V., Ragge N.K. (2024). Deletion upstream of MAB21L2 highlights the importance of evolutionarily conserved non-coding sequences for eye development. Nat Commun. 15(1):9245. Link
De Freitas J.T., Thakur V., LaPorte K.M., Thakur V.S., Flores B., Caicedo V., Ajaegbu C.G.E., Ingrasci G., Lipman Z.M., Zhang K., Qiu H., Malek T.R., Bedogni B. (2024). Notch1 blockade by a novel, selective anti-Notch1 neutralizing antibody improves immunotherapy efficacy in melanoma by promoting an inflamed TME. J Exp Clin Cancer Res. 43:295. Link
Dig B. Mahat., Nathaniel D. Tippens., Jorge D. Martin-Rufino., Sean K. Waterton., Jiayu Fu., Sarah E. Blatt & Phillip A. Sharp. (2024) Single-cell nascent RNA sequencing unveils coordinated global transcription. Nature. 631, 216–223. Link
Sung-Joon Park., Kenta Nakai. (2024) A computational approach for deciphering the interactions between proximal and distal gene regulators in GC B-cell response. NAR Genomics & Bioiformatics. Volume 6, issue 2. Link
Farrim M.I., Gomes A., Milenkovic D., Menezes R. (2024) Gene expression analysis reveals diabetes-related gene signatures. Hum Genomics 18, 16. Link
Eni-Aganga I., Lanaghan ZM., Ismail F., Korolkova O., Goodwin JS., Balasubramaniam M., Dash C., Pandhare J. (2024). KLF6 activates Sp1-mediated prolidase transcription during TGF-β1 signaling. J Biol Chem. 2024 300(2):105605. Link
Hasegawa K., Tamaki M., Sakamaki Y., Wakino S. (2024) Nmnat1 Deficiency Causes Mitoribosome Excess in Diabetic Nephropathy Mediated by Transcriptional Repressor HIC1. Int J Mol Sci. 25(12):6384. Link
Abrar M., Ali S., Hussain I., Khatoon H., Batool F., Ghazanfar S., Corcoran D., Kawakami Y., Abbasi AA. (2024). Cis-regulatory control of mammalian Trps1 gene expression. J Exp Zool B Mol Dev Evol. 342(2):85-100. Link
Scaramuzzo RT., Crucitta S., Del Re M., Cammalleri M., Bagnoli P., Dal Monte M., Pini A., Filippi L.. (2024) β3-adREnoceptor Analysis in CORD Blood of Neonates (β3 RECORD): Study Protocol of a Pilot Clinical Investigation. Life (Basel). 14(6):776. Link
Cene Skubic., Hana Trček., Petra Nassib., Tinkara Kreft., Andrew Walakira., Katka Pohar., Sara Petek., zadeja Režen., Alojz Ihan., Damjana Rozman. (2024) Knockouts of CYP51A1, DHCR24, or SC5D from cholesterol synthesis reveal pathways modulated by sterol intermediates. iScience, Volume 27, Issue 9, 110651. Link
Jamil M.A., Al-Rifai R., Nuesgen N., Altmüller J., Oldenburg J., El-Maarri O. (2024) The role of microRNAs in defining LSECs cellular identity and in regulating F8 gene expression. Front Genet. 15: 1302685 Link
Coatti G.C., Vaghela N., Gillurkar P., Leir S., Harris A. (2024) A promoter-dependent upstream activator augments CFTR expression in diverse epithelial cell types. Biochim Biophys Acta Gene Regul Mech. 1867(2):195031 Link
Satsu H., Gondo Y., Shimanaka H., Imae M., Murakami S., Watari K., Wakabayashi S., Park S.J., Nakai K., Shimizu M. (2022) Signaling Pathway of Taurine-Induced Upregulation of TXNIP. Metabolites. 12(7),636. Link
Deepti P., Pasha A., Kumbhakar D.V., Doneti R., Heena S.K., Bhanoth S., Poleboyina P.K., Yadala R., Anapurna S.D., Pawar S.C. (2022) Overexpression of Secreted Phosphoprotein 1 (SPP1) predicts poor survival in HPV positive cervical cancer. Gene. 824,146381. Link
Song Q., Bian Q., Liang T., Zhang Y., Zhang K. (2021) Identification of immune-related genes and susceptible population of pulmonary tuberculosis by constructing TF-miRNA-mRNA regulatory network. Tuberculosis (Edinb). 131,102139. LinkThompson B., Chen Y., Davidson E.A., Garcia-Milian R., Golla J.P., Apostolopoulos N., Orlicky D.J., Schey K., Thompson D.C., Vasiliou V. (2021) Impaired GSH biosynthesis disrupts eye development, lens morphogenesis and PAX6 function. Ocul Surf. 22,190-203. Link
Publications authored by the geneXplain team
Kisakol, B., Matveeva, A., Salvucci, M., Kel, A., McDonough, E., Ginty, F., Longley, D. B., & Prehn, J. H. M. (2024). Identification of unique rectal cancer-specific subtypes. British journal of cancer, 130(11), 1809–1818. Link
Kolpakov, F., Akberdin, I., Kiselev, I., Kolmykov, S., Kondrakhin, Y., Kulyashov, M., Kutumova, E., Pintus, S., Ryabova, A., Sharipov, R., Yevshin, I., Zhatchenko, S., & Kel, A. (2022). BioUML-towards a universal research platform. Nucleic Acids Res. 50(W1),W124–31. Link
Orekhov A.N., Sukhorukov V.N., Nikiforov N.G., Kubekina M.V., Sobenin I.A., Foxx K.K., Pintus S., Stegmaier P., Stelmashenko D., Kel A., Poznyak A.V., Wu W.K., Kasianov A.S., Makeev V.Y., Manabe I., Oishi Y. (2020) Signaling Pathways Potentially Responsible for Foam Cell Formation: Cholesterol Accumulation or Inflammatory Response-What is First? Int J Mol Sci. 21(8),2716. Link
Kel A., Boyarskikh U., Stegmaier P., Leskov L.S., Sokolov A.V., Yevshin I., Mandrik N., Stelmashenko D., Koschmann J., Kel-Margoulis O., Krull M., Martínez-Cardús A., Moran S., Esteller M., Kolpakov F., Filipenko M., Wingender E. (2019) Walking pathways with positive feedback loops reveal DNA methylation biomarkers of colorectal cancer. BMC Bioinformatics. 20(Suppl 4),119. Link
Boyarskikh, U., Pintus, S., Mandrik, N., Stelmashenko, D., Kiselev, I., Evshin, I., Sharipov, R., Stegmaier, P., Kolpakov, F., Filipenko, M., Kel, A. (2018) Computational master-regulator search reveals mTOR and PI3K pathways responsible for low sensitivity of NCI-H292 and A427 lung cancer cell lines to cytotoxic action of p53 activator Nutlin-3. BMC Med. Genomics 11(Suppl 1), 12. Link
Kel, A.E., Stegmaier, P., Valeev, T., Koschmann, J., Poroikov, V., Kel-Margoulis, O.V. and Wingender, E. (2016) Multi-omics “upstream analysis” of regulatory genomic regions helps identifying targets against methotrexate resistance of colon cancer. EuPA Open Proteomics 13, 1-13. Link
How to cite TRANSFAC
Matys V, Kel-Margoulis OV, Fricke E, Liebich I, Land S, Barre-Dirrie A, Reuter I, Chekmenev D, Krull M, Hornischer K, Voss N, Stegmaier P, Lewicki-Potapov B, Saxel H, Kel AE, Wingender E. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 2006;34(90001):D108-D110. Link